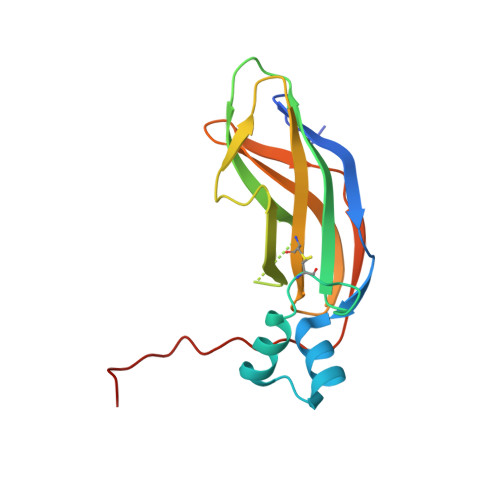
Structural conservation of the B subunit in the ammonia monooxygenase/particulate methane monooxygenase superfamily.
Lawton, T.J., Ham, J., Sun, T., Rosenzweig, A.C.(2014) Proteins 82: 2263-2267
- PubMed: 24523098 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.24535
- Primary Citation Related Structures:
4O65 - PubMed Abstract:
The ammonia monooxygenase (AMO)/particulate methane monooxygenase (pMMO) superfamily is a diverse group of membrane-bound enzymes of which only pMMO has been characterized on the molecular level. The pMMO active site is believed to reside in the soluble N-terminal region of the pmoB subunit. To understand the degree of structural conservation within this superfamily, the crystal structure of the corresponding domain of an archaeal amoB subunit from Nitrosocaldus yellowstonii has been determined to 1.8 Å resolution. The structure reveals a remarkable conservation of overall fold and copper binding site location as well as several notable differences that may have implications for function and stability.
- Departments of Molecular Biosciences and of Chemistry, Northwestern University, Evanston, Illinois, 60208.
Organizational Affiliation: