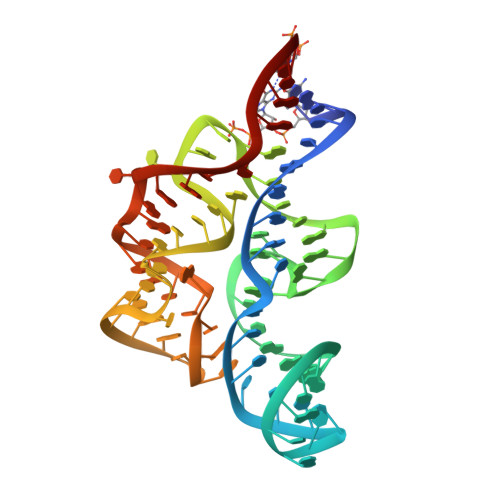
Validating Fragment-Based Drug Discovery for Biological RNAs: Lead Fragments Bind and Remodel the TPP Riboswitch Specifically.
Warner, K.D., Homan, P., Weeks, K.M., Smith, A.G., Abell, C., Ferre-D'Amare, A.R.(2014) Chem Biol 21: 591-595
- PubMed: 24768306 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2014.03.007
- Primary Citation Related Structures:
4NYA, 4NYB, 4NYC, 4NYD, 4NYG - PubMed Abstract:
Thiamine pyrophosphate (TPP) riboswitches regulate essential genes in bacteria by changing conformation upon binding intracellular TPP. Previous studies using fragment-based approaches identified small molecule "fragments" that bind this gene-regulatory mRNA domain. Crystallographic studies now show that, despite having micromolar Kds, four different fragments bind the TPP riboswitch site-specifically, occupying the pocket that recognizes the aminopyrimidine of TPP. Unexpectedly, the unoccupied site that would recognize the pyrophosphate of TPP rearranges into a structure distinct from that of the cognate complex. This idiosyncratic fragment-induced conformation, also characterized by small-angle X-ray scattering and chemical probing, represents a possible mechanism for adventitious ligand discrimination by the riboswitch, and suggests that off-pathway conformations of RNAs can be targeted for drug development. Our structures, together with previous screening studies, demonstrate the feasibility of fragment-based drug discovery against RNA targets.
- National Heart, Lung and Blood Institute, 50 South Drive, MSC 8012, Bethesda, MD 20892-8012, USA; Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge CB2 1EW, UK.
Organizational Affiliation: