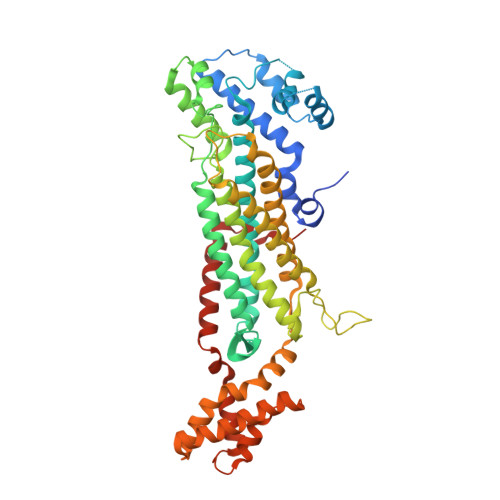
Structural and kinetic studies on adenylosuccinate lyase from Mycobacterium smegmatis and Mycobacterium tuberculosis provide new insights on the catalytic residues of the enzyme.
Banerjee, S., Agrawal, M.J., Mishra, D., Sharan, S., Balaram, H., Savithri, H.S., Murthy, M.R.(2014) FEBS J 281: 1642-1658
- PubMed: 24479855 Search on PubMed
- DOI: https://doi.org/10.1111/febs.12730
- Primary Citation Related Structures:
4NLE - PubMed Abstract:
Adenylosuccinate lyase (ASL), an enzyme involved in purine biosynthesis, has been recognized as a drug target against microbial infections. In the present study, ASL from Mycobacterium smegmatis (MsASL) and Mycobacterium tuberculosis (MtbASL) were cloned, purified and crystallized. The X-ray crystal structure of MsASL was determined at a resolution of 2.16 Å. It is the first report of an apo-ASL structure with a partially ordered active site C3 loop. Diffracting crystals of MtbASL could not be obtained and a model for its structure was derived using MsASL as a template. These structures suggest that His149 and either Lys285 or Ser279 of MsASL are the residues most likely to function as the catalytic acid and base, respectively. Most of the active site residues were found to be conserved, with the exception of Ser148 and Gly319 of MsASL. Ser148 is structurally equivalent to a threonine in most other ASLs. Gly319 is replaced by an arginine residue in most ASLs. The two enzymes were catalytically much less active compared to ASLs from other organisms. Arg319Gly substitution and reduced flexibility of the C3 loop might account for the low catalytic activity of mycobacterial ASLs. The low activity is consistent with the slow growth rate of Mycobacteria and their high GC containing genomes, as well as their dependence on other salvage pathways for the supply of purine nucleotides. purB and purB bind by x-ray crystallography (View interaction).
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: