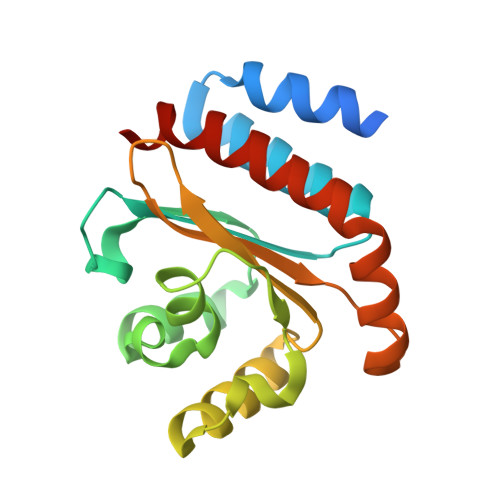
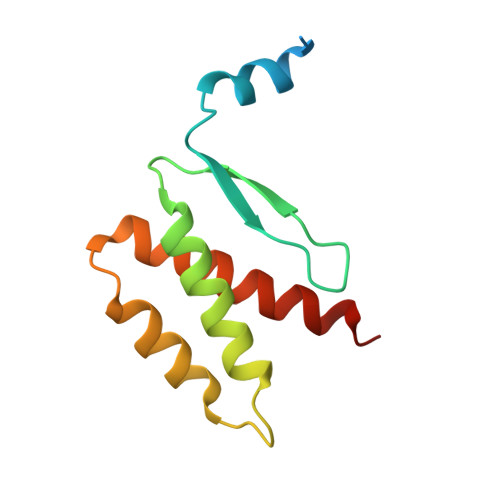
QsIA disrupts LasR dimerization in antiactivation of bacterial quorum sensing
Fan, H., Dong, Y., Wu, D.H., Bowler, M.W., Zhang, L., Song, H.(2013) Proc Natl Acad Sci U S A 110: 20765-20770
- PubMed: 24319092 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1314415110
- Primary Citation Related Structures:
4NG2 - PubMed Abstract:
The human pathogen Pseudomonas aeruginosa coordinates the expression of virulence factors by using quorum sensing (QS), a signaling cascade triggered by the QS signal molecule and its receptor, a member of the LuxR family of QS transcriptional factors (LasR). The QS threshold and response in P. aeruginosa is defined by a QS LasR-specific antiactivator (QslA), which binds to LasR and prevents it from binding to its target promoter. However, how QslA binds to LasR and regulates its DNA binding activity in QS remains elusive. Here we report the crystal structure of QslA in complex with the N-terminal ligand binding domain of LasR. QsIA exists as a functional dimer to interact with the LasR ligand binding domain. Further analysis shows that QsIA binding occupies the LasR dimerization interface and consequently disrupts LasR dimerization, thereby preventing LasR from binding to its target DNA and disturbing normal QS. Our findings provide a structural model for understanding the QslA-mediated antiactivation mechanism in QS through protein-protein interaction.
- Institute of Molecular and Cell Biology, Singapore 138673.
Organizational Affiliation: