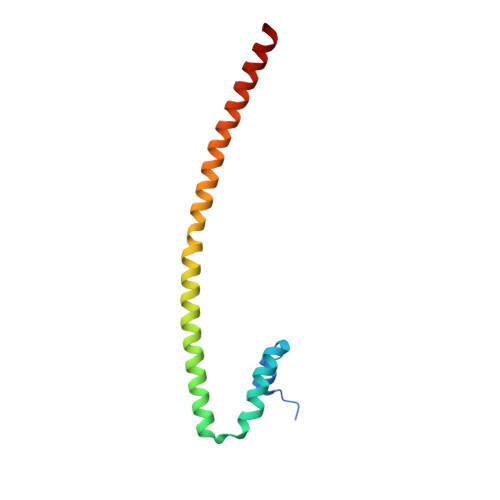
Crystal structure of the nipah virus phosphoprotein tetramerization domain.
Bruhn, J.F., Barnett, K.C., Bibby, J., Thomas, J.M., Keegan, R.M., Rigden, D.J., Bornholdt, Z.A., Saphire, E.O.(2014) J Virol 88: 758-762
- PubMed: 24155387 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.02294-13
- Primary Citation Related Structures:
4N5B - PubMed Abstract:
The Nipah virus phosphoprotein (P) is multimeric and tethers the viral polymerase to the nucleocapsid. We present the crystal structure of the multimerization domain of Nipah virus P: a long, parallel, tetrameric, coiled coil with a small, α-helical cap structure. Across the paramyxoviruses, these domains share little sequence identity yet are similar in length and structural organization, suggesting a common requirement for scaffolding or spatial organization of the functions of P in the virus life cycle.
- Department of Immunology and Microbial Science, The Scripps Research Institute, La Jolla, California, USA.
Organizational Affiliation: