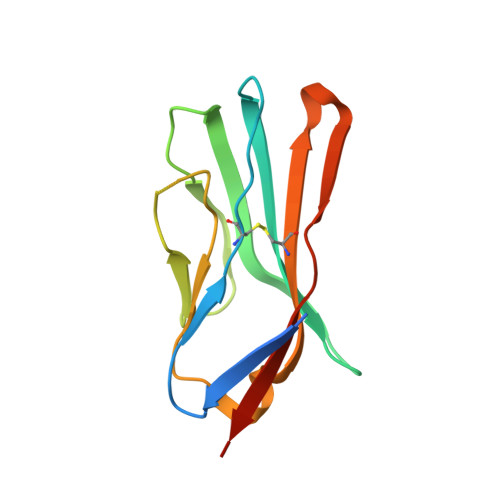
Crystallographic insights into sodium-channel modulation by the beta 4 subunit.
Gilchrist, J., Das, S., Van Petegem, F., Bosmans, F.(2013) Proc Natl Acad Sci U S A 110: E5016-E5024
- PubMed: 24297919 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1314557110
- Primary Citation Related Structures:
4MZ2, 4MZ3 - PubMed Abstract:
Voltage-gated sodium (Nav) channels are embedded in a multicomponent membrane signaling complex that plays a crucial role in cellular excitability. Although the mechanism remains unclear, β-subunits modify Nav channel function and cause debilitating disorders when mutated. While investigating whether β-subunits also influence ligand interactions, we found that β4 dramatically alters toxin binding to Nav1.2. To explore these observations further, we solved the crystal structure of the extracellular β4 domain and identified (58)Cys as an exposed residue that, when mutated, eliminates the influence of β4 on toxin pharmacology. Moreover, our results suggest the presence of a docking site that is maintained by a cysteine bridge buried within the hydrophobic core of β4. Disrupting this bridge by introducing a β1 mutation implicated in epilepsy repositions the (58)Cys-containing loop and disrupts β4 modulation of Nav1.2. Overall, the principles emerging from this work (i) help explain tissue-dependent variations in Nav channel pharmacology; (ii) enable the mechanistic interpretation of β-subunit-related disorders; and (iii) provide insights in designing molecules capable of correcting aberrant β-subunit behavior.
- Department of Physiology and Solomon H. Snyder Department of Neuroscience, The Johns Hopkins University School of Medicine, Baltimore, MD 21205.
Organizational Affiliation: