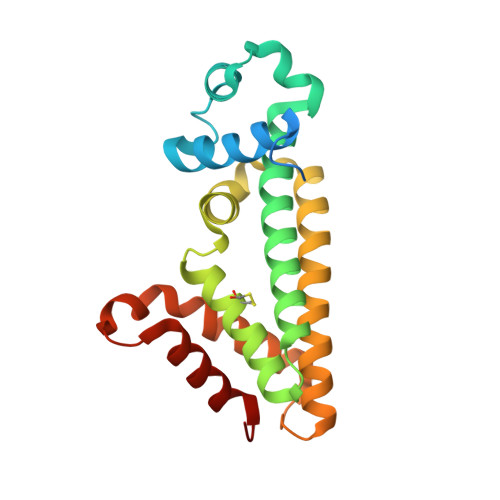
Pfit is a structurally novel Crohn's disease-associated superantigen.
Liu, L., Chen, H., Brecher, M.B., Li, Z., Wei, B., Nandi, B., Zhang, J., Ling, H., Winslow, G., Braun, J., Li, H.(2013) PLoS Pathog 9: e1003837-e1003837
- PubMed: 24385909 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1003837
- Primary Citation Related Structures:
4MO7, 4MXM - PubMed Abstract:
T cell responses to enteric bacteria are important in inflammatory bowel disease. I2, encoded by the pfiT gene of Pseudomonas fluorescens, is a T-cell superantigen associated with human Crohn's disease. Here we report the crystal structure of pfiT at 1.7Å resolution and provide a functional analysis of the interaction of pfiT and its homolog, PA2885, with human class II MHC. Both pfiT and PA2885 bound to mammalian cells and stimulated the proliferation of human lymphocytes. This binding was greatly inhibited by anti-class II MHC HLA-DR antibodies, and to a lesser extent, by anti HLA-DQ and DP antibodies, indicating that the binding was class II MHC-specific. GST-pfiT efficiently precipitated both endogenous and in vitro purified recombinant HLA-DR1 molecules, indicating that pfiT directly interacted with HLA-DR1. Competition studies revealed that pfiT and the superantigen Mycoplasma arthritidis mitogen (MAM) competed for binding to HLA-DR, indicating that their binding sites overlap. Structural analyses established that pfiT belongs to the TetR-family of DNA-binding transcription regulators. The distinct structure of pfiT indicates that it represents a new family of T cell superantigens.
- Wadsworth Center, New York State Department of Health, Albany, New York, United States of America.
Organizational Affiliation: