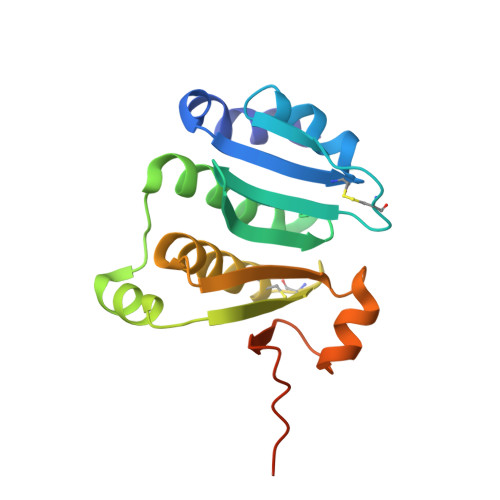
Sulfolobus solfataricus thiol redox puzzle: characterization of an atypical protein disulfide oxidoreductase.
Limauro, D., De Simone, G., Pirone, L., Bartolucci, S., D'Ambrosio, K., Pedone, E.(2014) Extremophiles 18: 219-228
- PubMed: 24306780 Search on PubMed
- DOI: https://doi.org/10.1007/s00792-013-0607-8
- Primary Citation Related Structures:
4MNN - PubMed Abstract:
Protein disulfide oxidoreductases (PDOs) are proteins involved in disulfide bond formation playing a crucial role in adaptation to extreme environment. This paper reports the functional and structural characterization of Sso1120, a PDO from the hyperthermophilic archaeon Sulfolobus solfataricus. The protein was expressed in Escherichia coli and purified to homogeneity. The functional characterization showed that the enzyme has reductase activity, as tested by insulin assay, but differently from the other PDOs, it does not present isomerase activity. In addition it is able to form a redox couple with the thioredoxin reductase that could be used in undiscovered pathways. The protein revealed a melting point of around 90 °C in CD spectroscopy-monitored thermal denaturation and high denaturant resistance. The X-ray crystallographic structure was solved at 1.80 Å resolution, showing differences with respect to other PDOs and an unexpected similarity with the N-terminal domain of the alkyl hydroperoxide reductase F component from Salmonella typhimurium. On the basis of the reported data and of bioinformatics and phylogenetic analyses, a possible involvement of this atypical PDO in a new antioxidant system of S. solfataricus has been proposed.
- Dipartimento di Biologia, Università degli Studi di Napoli ''Federico II'', Complesso Universitario Monte S. Angelo, Via Cinthia, 80126, Naples, Italy.
Organizational Affiliation: