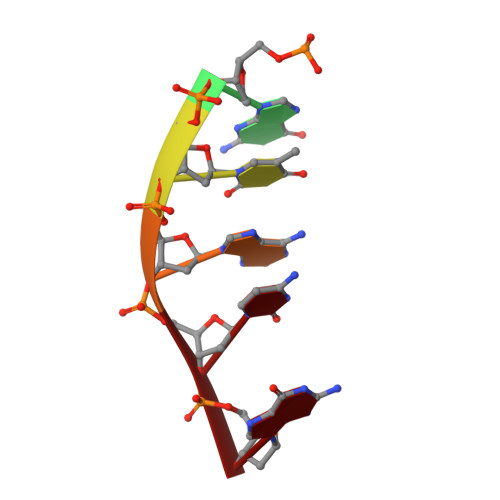
Intercalative DNA binding of the marine anticancer drug variolin B.
Canals, A., Arribas-Bosacoma, R., Albericio, F., Alvarez, M., Aymami, J., Coll, M.(2017) Sci Rep 7: 39680-39680
- PubMed: 28051169 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep39680
- Primary Citation Related Structures:
4MNB - PubMed Abstract:
Variolin B is a rare marine alkaloid that showed promising anti-cancer activity soon after its isolation. It acts as a cyclin-dependent kinase inhibitor, although the precise mechanism through which it exerts the cytotoxic effects is still unknown. The crystal structure of a variolin B bound to a DNA forming a pseudo-Holliday junction shows that this compound can also contribute, through intercalative binding, to either the formation or stabilization of multi-stranded DNA forms.
- Institute for Research in Biomedicine (IRB Barcelona), The Barcelona Institute of Science and Technology, Baldiri Reixac 10, 08028 Barcelona, Spain.
Organizational Affiliation: