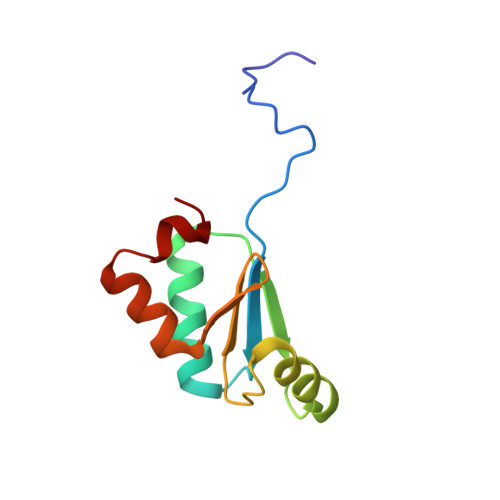
Utility of Synechocystis sp. PCC 6803 glutaredoxin A as a platform to study high-resolution mutagenesis of proteins.
Knaff, D.B., Sutton, R.B.(2013) Front Plant Sci 4: 461-461
- PubMed: 24298277 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fpls.2013.00461
- Primary Citation Related Structures:
4MJA, 4MJB, 4MJC, 4MJE - PubMed Abstract:
Glutaredoxin from the cyanobacterium Synechocystis sp. PCC 6803 is a small protein, containing only 88 amino acids, that participates in a large number of redox reactions, serving both as an electron donor for enzyme-catalyzed reductions and as a regulator of diverse metabolic pathways. The crystal structures of glutaredoxins from several species have been solved, including the glutaredoxin A isoform from the cyanobacterium Synechocystis sp. PCC 6803. We have utilized the small size of Synechocystis glutaredoxin A and its propensity to form protein crystals that diffract to high resolution to explore a long-standing question in biochemistry; i.e., what are the effects of mutations on protein structure and function? Taking advantage of these properties, we have initiated a long-term educational project that would examine the structural and biochemical changes in glutaredoxin as a function of single-point mutational replacements. Here, we report some of the mutational effects that we have observed to date.
- Department of Chemistry and Biochemistry, Texas Tech University Lubbock, TX, USA ; Center for Biotechnology and Genomics, Texas Tech University Lubbock, TX, USA.
Organizational Affiliation: