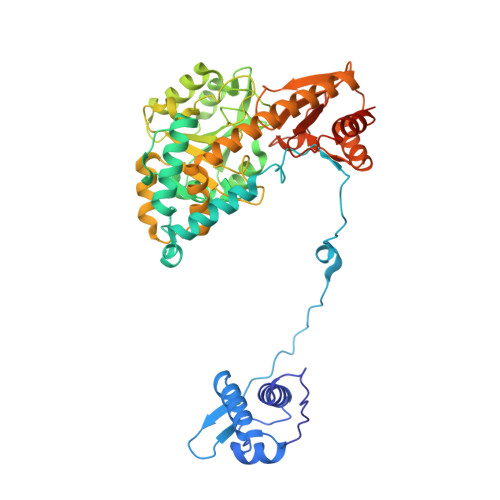
Crystal structure of Bacillus subtilis GabR, an autorepressor and transcriptional activator of gabT.
Edayathumangalam, R., Wu, R., Garcia, R., Wang, Y., Wang, W., Kreinbring, C.A., Bach, A., Liao, J., Stone, T.A., Terwilliger, T.C., Hoang, Q.Q., Belitsky, B.R., Petsko, G.A., Ringe, D., Liu, D.(2013) Proc Natl Acad Sci U S A 110: 17820-17825
- PubMed: 24127574 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1315887110
- Primary Citation Related Structures:
4MGR, 4N0B - PubMed Abstract:
Bacillus subtilis GabR is a transcription factor that regulates gamma-aminobutyric acid (GABA) metabolism. GabR is a member of the understudied MocR/GabR subfamily of the GntR family of transcription regulators. A typical MocR/GabR-type regulator is a chimeric protein containing a short N-terminal helix-turn-helix DNA-binding domain and a long C-terminal pyridoxal 5'-phosphate (PLP)-binding putative aminotransferase domain. In the presence of PLP and GABA, GabR activates the gabTD operon, which allows the bacterium to use GABA as nitrogen and carbon sources. GabR binds to its own promoter and represses gabR transcription in the absence of GABA. Here, we report two crystal structures of full-length GabR from B. subtilis: a 2.7-Å structure of GabR with PLP bound and the 2.55-Å apo structure of GabR without PLP. The quaternary structure of GabR is a head-to-tail domain-swap homodimer. Each monomer comprises two domains: an N-terminal winged-helix DNA-binding domain and a C-terminal PLP-binding type I aminotransferase-like domain. The winged-helix domain contains putative DNA-binding residues conserved in other GntR-type regulators. Together with sedimentation velocity and fluorescence polarization assays, the crystal structure of GabR provides insights into DNA binding by GabR at the gabR and gabT promoters. The absence of GabR-mediated aminotransferase activity in the presence of GABA and PLP, and the presence of an active site configuration that is incompatible with stabilization of the GABA external aldimine suggest that a GabR aminotransferase-like activity involving GABA and PLP is not essential to its primary function as a transcription regulator.
- Center for Neurologic Diseases, Harvard Medical School and Brigham and Women's Hospital, Boston, MA 02115.
Organizational Affiliation: