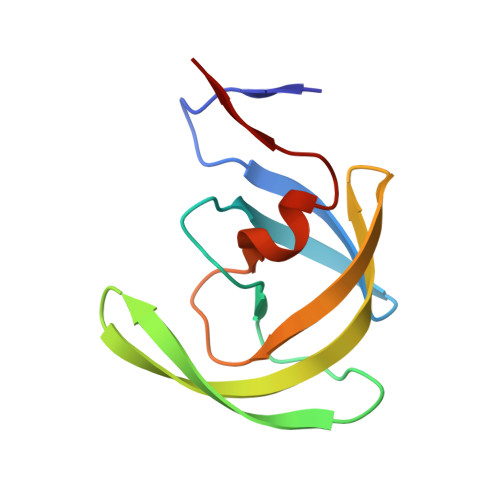
Structural Optimization of Cyclic Sulfonamide based Novel HIV-1 Protease Inhibitors to Pico Molar Affinities guided by X-ray Crystallographic Analysis
Ganguly, A.K., Alluri, S.S., Wang, C.H., Antropow, A., White, A., Caroccia, D., Biswas, D., Kang, E., Zhang, L.K., Carroll, S.S., Burlein, C., Fay, J., Orth, P., Strickland, C.(2014) Tetrahedron