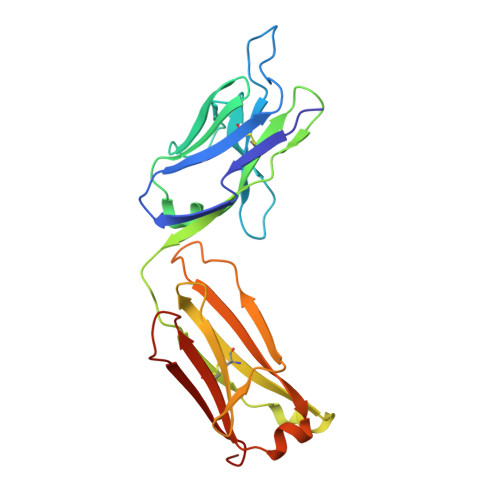
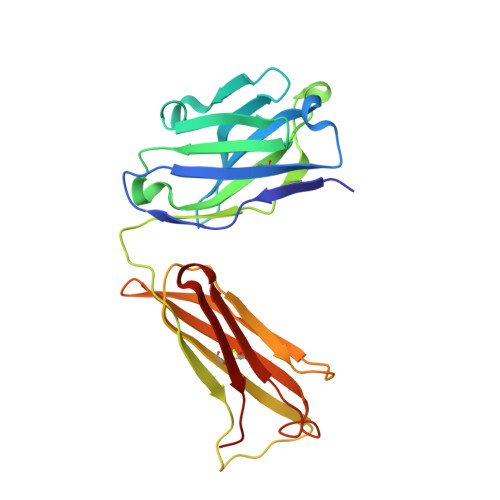
Crystal structure determination of anti-DNA Fab A52.
Stanfield, R.L., Eilat, D.(2014) Proteins 82: 1674-1678
- PubMed: 24449198 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24514
- Primary Citation Related Structures:
4M61 - PubMed Abstract:
A52 is a murine monoclonal antibody isolated from autoimmune New Zealand Black/New Zealand White F1 mice that recognizes single and double stranded DNA. This mouse strain spontaneously develops systemic lupus erythematosus-like symptoms and has served as a model for that disease for many years. The 1.62 Å crystal structure of the A52 Fab fragment reveals an H3 complementarity determining region with four closely spaced arginine residues, creating a positively charged surface to accommodate bound DNA.
- The Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, California, 92037.
Organizational Affiliation: