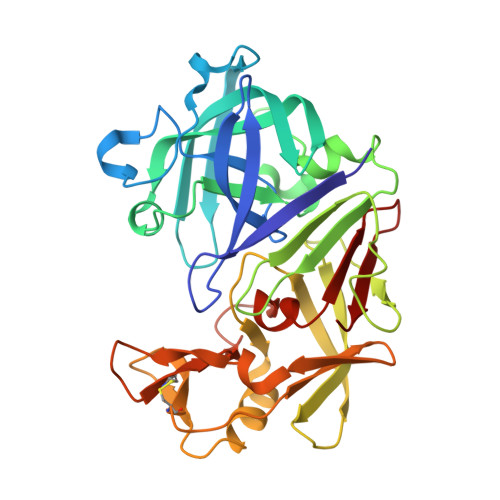
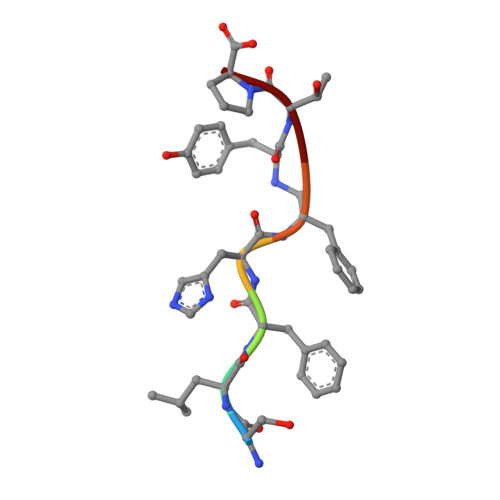
The structure of endothiapepsin complexed with a Phe-Tyr reduced-bond inhibitor at 1.35 angstrom resolution.
Guo, J., Cooper, J.B., Wood, S.P.(2014) Acta Crystallogr D Biol Crystallogr 70: 30-33
- PubMed: 24419612 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X13032974
- Primary Citation Related Structures:
4LP9 - PubMed Abstract:
Endothiapepsin is a typical member of the aspartic proteinase family. The catalytic mechanism of this family is attributed to two conserved catalytic aspartate residues, which coordinate the hydrolysis of a peptide bond. An oligopeptide inhibitor (IC50 = 0.62 µM) based on a reduced-bond transition-state inhibitor of mucorpepsin was co-crystallized with endothiapepsin and the crystal structure of the enzyme-inhibitor complex was determined at 1.35 Å resolution. A total of 12 hydrogen bonds between the inhibitor and the active-site residues were identified. The resulting structure demonstrates a number of novel subsite interactions in the active-site cleft.
- Laboratory of Protein Crystallography, Centre for Amyloidosis and Acute Phase Proteins, UCL Division of Medicine (Royal Free Campus), Rowland Hill Street, London NW3 2PF, England.
Organizational Affiliation: