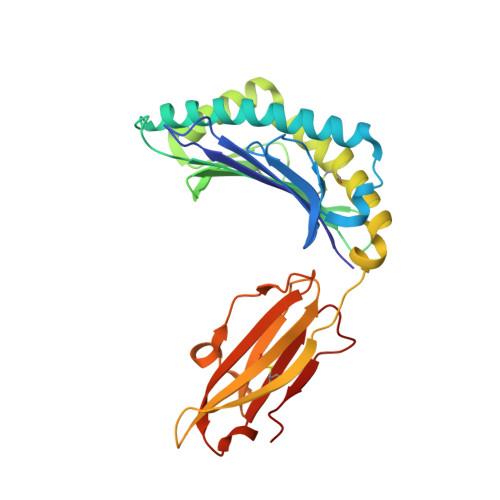
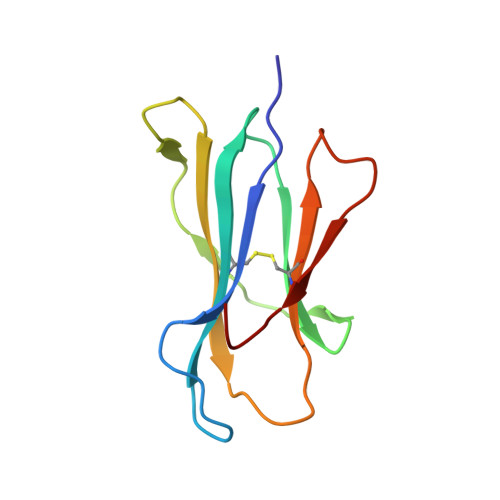
Peptide-dependent conformational fluctuation determines the stability of the human leukocyte antigen class I complex.
Yanaka, S., Ueno, T., Shi, Y., Qi, J., Gao, G.F., Tsumoto, K., Sugase, K.(2014) J Biological Chem 289: 24680-24690
- PubMed: 25028510 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.566174
- Primary Citation Related Structures:
4LNR - PubMed Abstract:
In immune-mediated control of pathogens, human leukocyte antigen (HLA) class I presents various antigenic peptides to CD8(+) T-cells. Long-lived peptide presentation is important for efficient antigen-specific T-cell activation. Presentation time depends on the peptide sequence and the stability of the peptide-HLA complex (pHLA). However, the determinant of peptide-dependent pHLA stability remains elusive. Here, to reveal the pHLA stabilization mechanism, we examined the crystal structures of an HLA class I allomorph in complex with HIV-derived peptides and evaluated site-specific conformational fluctuations using NMR. Although the crystal structures of various pHLAs were almost identical independent of the peptides, fluctuation analyses identified a peptide-dependent minor state that would be more tightly packed toward the peptide. The minor population correlated well with the thermostability and cell surface presentation of pHLA, indicating that this newly identified minor state is important for stabilizing the pHLA and facilitating T-cell recognition.
- From the Department of Medical Genome Sciences, Graduate School of Frontier Sciences, The University of Tokyo, Tokyo 277-8562, Japan.
Organizational Affiliation: