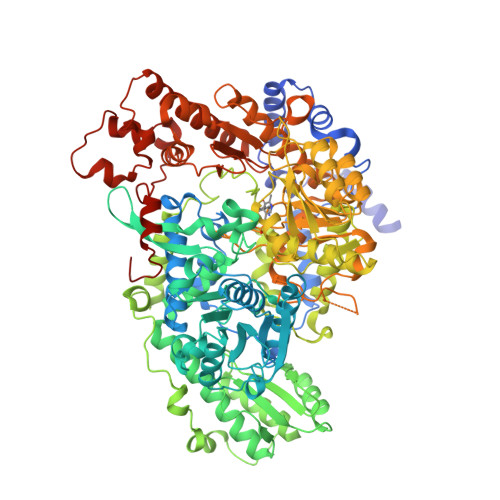
Structure of the Homodimeric Glycine Decarboxylase P-protein from Synechocystis sp. PCC 6803 Suggests a Mechanism for Redox Regulation.
Hasse, D., Andersson, E., Carlsson, G., Masloboy, A., Hagemann, M., Bauwe, H., Andersson, I.(2013) J Biological Chem 288: 35333-35345
- PubMed: 24121504 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.509976
- Primary Citation Related Structures:
4LGL, 4LHC, 4LHD - PubMed Abstract:
Glycine decarboxylase, or P-protein, is a pyridoxal 5'-phosphate (PLP)-dependent enzyme in one-carbon metabolism of all organisms, in the glycine and serine catabolism of vertebrates, and in the photorespiratory pathway of oxygenic phototrophs. P-protein from the cyanobacterium Synechocystis sp. PCC 6803 is an α2 homodimer with high homology to eukaryotic P-proteins. The crystal structure of the apoenzyme shows the C terminus locked in a closed conformation by a disulfide bond between Cys(972) in the C terminus and Cys(353) located in the active site. The presence of the disulfide bridge isolates the active site from solvent and hinders the binding of PLP and glycine in the active site. Variants produced by substitution of Cys(972) and Cys(353) by Ser using site-directed mutagenesis have distinctly lower specific activities, supporting the crucial role of these highly conserved redox-sensitive amino acid residues for P-protein activity. Reduction of the 353-972 disulfide releases the C terminus and allows access to the active site. PLP and the substrate glycine bind in the active site of this reduced enzyme and appear to cause further conformational changes involving a flexible surface loop. The observation of the disulfide bond that acts to stabilize the closed form suggests a molecular mechanism for the redox-dependent activation of glycine decarboxylase observed earlier.
- From the Department of Cell and Molecular Biology, Uppsala University, S-751 24 Uppsala, Sweden.
Organizational Affiliation: