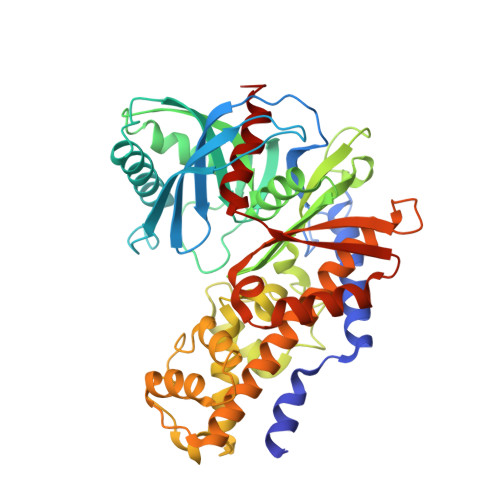
Pyrimidone-based series of glucokinase activators with alternative donor-acceptor motif.
Filipski, K.J., Guzman-Perez, A., Bian, J., Perreault, C., Aspnes, G.E., Didiuk, M.T., Dow, R.L., Hank, R.F., Jones, C.S., Maguire, R.J., Tu, M., Zeng, D., Liu, S., Knafels, J.D., Litchfield, J., Atkinson, K., Derksen, D.R., Bourbonais, F., Gajiwala, K.S., Hickey, M., Johnson, T.O., Humphries, P.S., Pfefferkorn, J.A.(2013) Bioorg Med Chem Lett 23: 4571-4578
- PubMed: 23831135 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2013.06.036
- Primary Citation Related Structures:
4L3Q - PubMed Abstract:
Glucokinase activators are a class of experimental agents under investigation as a therapy for Type 2 diabetes mellitus. An X-ray crystal structure of a modestly potent agent revealed the potential to substitute the common heterocyclic amide donor-acceptor motif for a pyridone moiety. We have successfully demonstrated that both pyridone and pyrimidone heterocycles can be used as a potent donor-acceptor substituent. Several sub-micromolar analogs that possess the desired partial activator profile were synthesized and characterized. Unfortunately, the most potent activators suffered from sub-optimal pharmacokinetic properties. Nonetheless, these donor-acceptor motifs may find utility in other glucokinase activator series or beyond.
- Cardiovascular, Metabolic, and Endocrine Diseases Chemistry, Pfizer Worldwide Research and Development, 620 Memorial Dr, Cambridge, MA 02139, USA. kevin.filipski@pfizer.com
Organizational Affiliation: