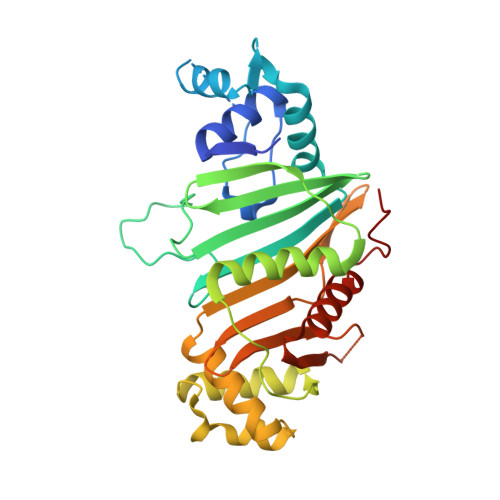
Structure of the Mycobacterium tuberculosis type VII secretion system chaperone EspG5 in complex with PE25-PPE41 dimer.
Korotkova, N., Freire, D., Phan, T.H., Ummels, R., Creekmore, C.C., Evans, T.J., Wilmanns, M., Bitter, W., Parret, A.H., Houben, E.N., Korotkov, K.V.(2014) Mol Microbiol 94: 367-382
- PubMed: 25155747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/mmi.12770
- Primary Citation Related Structures:
4KXR - PubMed Abstract:
The growth or virulence of Mycobacterium tuberculosis bacilli depends on homologous type VII secretion systems, ESX-1, ESX-3 and ESX-5, which export a number of protein effectors across membranes to the bacterial surface and environment. PE and PPE proteins represent two large families of highly polymorphic proteins that are secreted by these ESX systems. Recently, it was shown that these proteins require system-specific cytoplasmic chaperones for secretion. Here, we report the crystal structure of M. tuberculosis ESX-5-secreted PE25-PPE41 heterodimer in complex with the cytoplasmic chaperone EspG(5). EspG(5) represents a novel fold that is unrelated to previously characterized secretion chaperones. Functional analysis of the EspG(5) -binding region uncovered a hydrophobic patch on PPE41 that promotes dimer aggregation, and the chaperone effectively abolishes this process. We show that PPE41 contains a characteristic chaperone-binding sequence, the hh motif, which is highly conserved among ESX-1-, ESX-3- and ESX-5-specific PPE proteins. Disrupting the interaction between EspG(5) and three different PPE target proteins by introducing different point mutations generally affected protein secretion. We further demonstrate that the EspG(5) chaperone plays an important role in the ESX secretion mechanism by keeping aggregation-prone PE-PPE proteins in their soluble state.
- Department of Molecular & Cellular Biochemistry, University of Kentucky, Lexington, KY, 40536, USA; Center for Structural Biology, University of Kentucky, Lexington, KY, 40536, USA.
Organizational Affiliation: