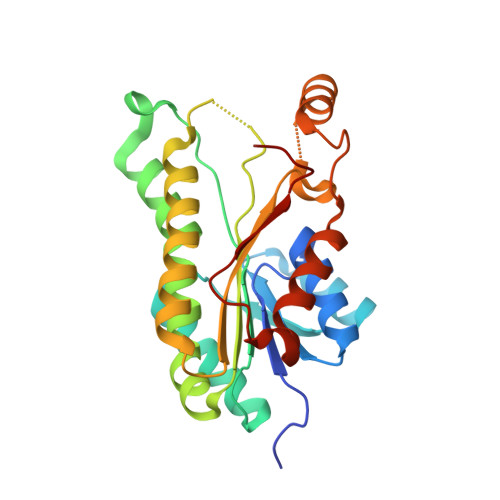
Crystal structure of acetoacetyl-CoA reductase from Rickettsia felis.
Rodarte, J.V., Abendroth, J., Edwards, T.E., Lorimer, D.D., Staker, B.L., Zhang, S., Myler, P.J., McLaughlin, K.J.(2021) Acta Crystallogr F Struct Biol Commun 77: 54-60
- PubMed: 33620038 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X21001497
- Primary Citation Related Structures:
4KMS, 7MI0 - PubMed Abstract:
Rickettsia felis, a Gram-negative bacterium that causes spotted fever, is of increasing interest as an emerging human pathogen. R. felis and several other Rickettsia strains are classed as National Institute of Allergy and Infectious Diseases priority pathogens. In recent years, R. felis has been shown to be adaptable to a wide range of hosts, and many fevers of unknown origin are now being attributed to this infectious agent. Here, the structure of acetoacetyl-CoA reductase from R. felis is reported at a resolution of 2.0 Å. While R. felis acetoacetyl-CoA reductase shares less than 50% sequence identity with its closest homologs, it adopts a fold common to other short-chain dehydrogenase/reductase (SDR) family members, such as the fatty-acid synthesis II enzyme FabG from the prominent pathogens Staphylococcus aureus and Bacillus anthracis. Continued characterization of the Rickettsia proteome may prove to be an effective means of finding new avenues of treatment through comparative structural studies.
- Department of Chemistry, Vassar College, 124 Raymond Avenue, Poughkeepsie, New York, USA.
Organizational Affiliation: