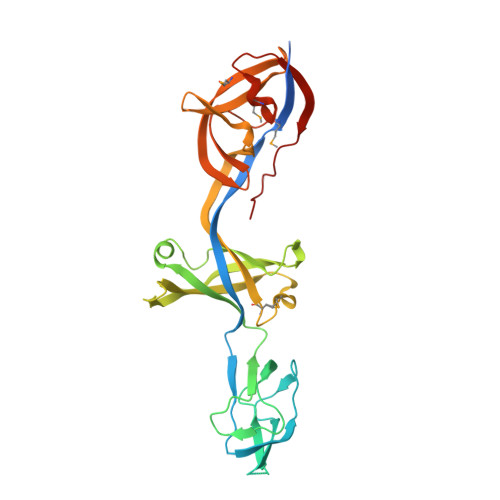
Structure of an atypical periplasmic adaptor from a multidrug efflux pump of the spirochete Borrelia burgdorferi.
Greene, N.P., Hinchliffe, P., Crow, A., Ababou, A., Hughes, C., Koronakis, V.(2013) FEBS Lett 587: 2984-2988
- PubMed: 23851070 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.febslet.2013.06.056
- Primary Citation Related Structures:
4KKS, 4KKT, 4KKU - PubMed Abstract:
Periplasmic adaptor proteins are essential components of bacterial tripartite multidrug efflux pumps. Here we report the 2.35 Å resolution crystal structure of the BesA adaptor from the spirochete Borrelia burgdorferi solved using selenomethionine derivatized protein. BesA shows the archetypal linear, flexible, multi-domain architecture evident among proteobacteria and retains the lipoyl, β-barrel and membrane-proximal domains that interact with the periplasmic domains of the inner membrane transporter. However, it lacks the α-hairpin domain shown to establish extensive coiled-coil interactions with the periplasmic entrance helices of the outer membrane-anchored TolC exit duct. This has implications for the modelling of assembled tripartite efflux pumps.
- Department of Pathology, University of Cambridge, Tennis Court Road, Cambridge CB2 1QP, UK.
Organizational Affiliation: