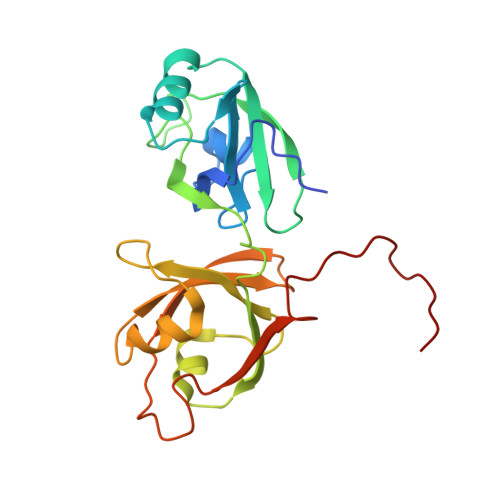
Structural insight into Golgi membrane stacking by GRASP65 and GRASP55 proteins
Feng, Y., Yu, W., Li, X., Lin, S., Zhou, Y., Hu, J., Liu, X.(2013) J Biological Chem 288: 28418-28427
- PubMed: 23940043 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.478024
- Primary Citation Related Structures:
4KFV, 4KFW - PubMed Abstract:
The stacking of Golgi cisternae involves GRASP65 and GRASP55. The oligomerization of the N-terminal GRASP domain of these proteins, which consists of two tandem PDZ domains, is required to tether the Golgi membranes. However, the molecular basis for GRASP assembly is unclear. Here, we determined the crystal structures of the GRASP domain of GRASP65 and GRASP55. The structures reveal similar homotypic interactions: the GRASP domain forms a dimer in which the peptide-binding pockets of the two neighboring PDZ2 domains face each other, and the dimers are further connected by the C-terminal tail of one GRASP domain inserting into the binding pocket of the PDZ1 domain in another dimer. Biochemical analysis suggests that both types of contacts are relatively weak but are needed in combination for GRASP-mediated Golgi stacking. Our results unveil a novel mode of membrane tethering by GRASP proteins and provide insight into the mechanism of Golgi stacking.
- From the Department of Biochemistry and Molecular Biology, College of Life Sciences, and State Key Laboratory of Medicinal Chemical Biology and.
Organizational Affiliation: