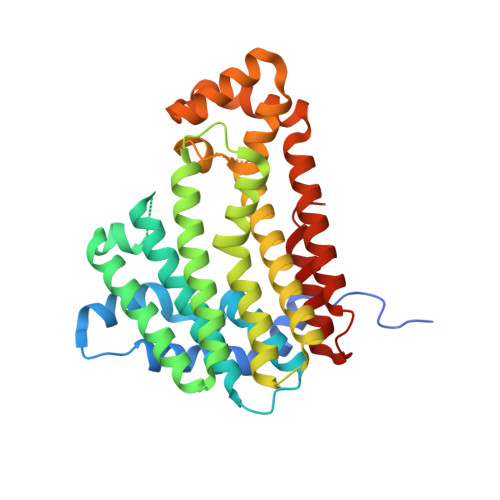
CRYSTAL STRUCTURE OF ISOPRENOID SYNTHASE PATL_3739 FROM Pseudoalteromonas atlantica
Patskovsky, Y., Toro, R., Bhosle, R., Hillerich, B., Seidel, R.D., Washington, E., Scott Glenn, A., Chowdhury, S., Evans, B., Hammonds, J., Zencheck, W.D., Imker, H.J., Al Obaidi, N., Stead, M., Love, J., Poulter, C.D., Gerlt, J.A., Almo, S.C.To be published.