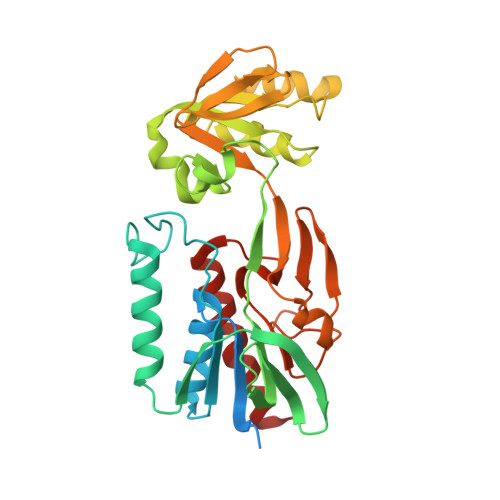
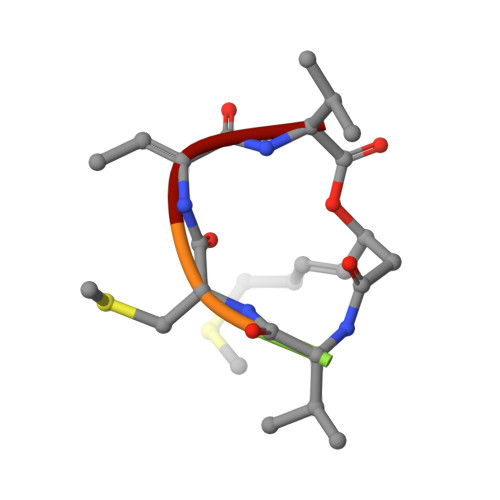
The structural basis of an NADP+-independent dithiol oxidase in FK228 biosynthesis.
Li, J., Wang, C., Zhang, Z.M., Cheng, Y.Q., Zhou, J.(2014) Sci Rep 4: 4145-4145
- PubMed: 24553401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep04145
- Primary Citation Related Structures:
4JN9, 4JNA - PubMed Abstract:
The disulfide bond is unusual in natural products and critical for thermal stability, cell permeability and bioactivity. DepH from Chromobacterium violaceum No. 968 is an FAD-dependent enzyme responsible for catalyzing the disulfide bond formation of FK228, an anticancer prodrug approved for the treatment of cutaneous T-cell lymphoma. Here we report the crystal structures of DepH and DepH complexed with a substrate analogue S,S'-dimethyl FK228 at 1.82 Å and 2.00 Å, respectively. Structural and biochemical analyses revealed that DepH, in contrast to the well characterized low molecular weight thioredoxin reductases (LMW TrxRs), is an NADP(+)-independent dithiol oxidase. DepH not only lacks a conserved GGGDXAXE motif necessary for NADP(+) binding in the canonical LMW TrxRs, but also contains a 11-residue sequence which physically impedes the binding of NADP(+). These observations explain the difference between NADP(+)-independent small molecule dithiol oxidases and NADP(+)-dependent thioredoxin reductases and provide insights for understanding the catalytic mechanism of dithiol oxidases involved in natural product biosynthesis.
- 1] State Key Laboratory of Bio-organic and Natural Products Chemistry, Shanghai Institute of Organic Chemistry, Chinese Academy of Sciences, Shanghai 200032, China [2].
Organizational Affiliation: