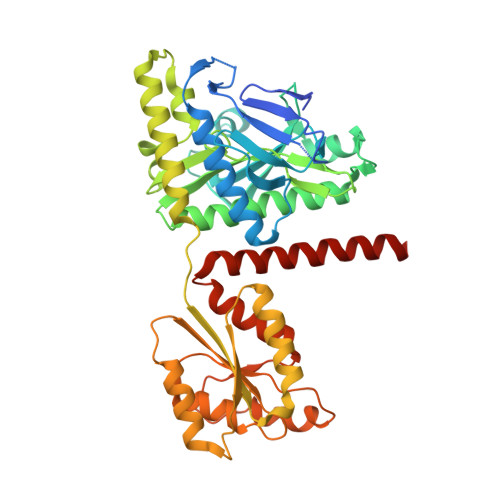
Crystal structure of the invertebrate bifunctional purine biosynthesis enzyme PAICS at 2.8 angstrom resolution.
Taschner, M., Basquin, J., Benda, C., Lorentzen, E.(2013) Proteins 81: 1473-1478
- PubMed: 23553965 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24296
- Primary Citation Related Structures:
4JA0 - PubMed Abstract:
Two important steps of the de novo purine biosynthesis pathway are catalyzed by the 5-aminoimidazole ribonucleotide carboxylase and the 4-(N-succinylcarboxamide)-5-aminoimidazole ribonucleotide synthetase enzymes. In most eukaryotic organisms, these two activities are present in the bifunctional enzyme complex known as PAICS. We have determined the 2.8-Å resolution crystal structure of the 350-kDa invertebrate PAICS from insect cells (Trichoplusia ni) using single-wavelength anomalous dispersion methods. Comparison of insect PAICS to human and prokaryotic homologs provides insights into substrate binding and reveals a highly conserved enzymatic framework across divergent species.
- Max-Planck-Institute of Biochemistry, Department of Structural Cell Biology, Martinsried, Germany.
Organizational Affiliation: