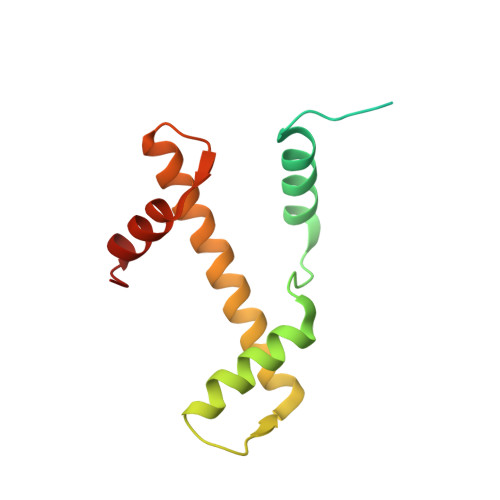
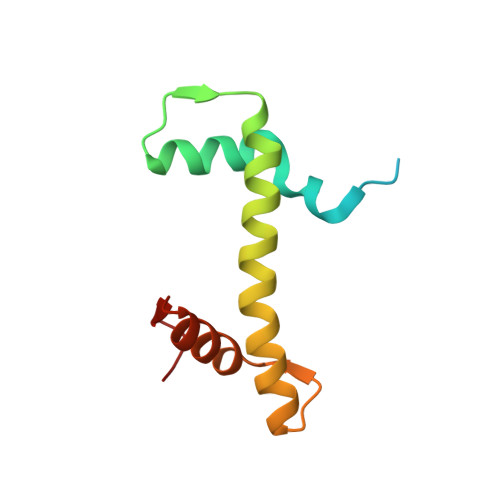
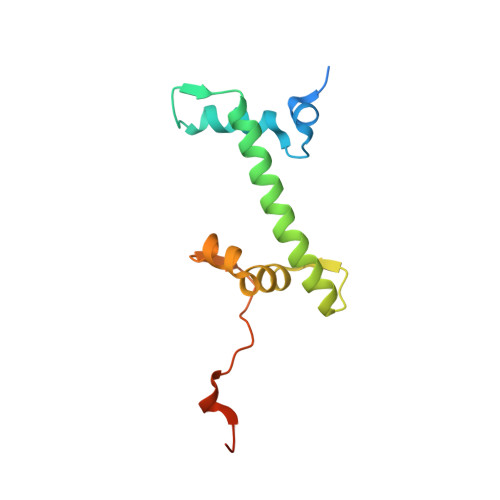
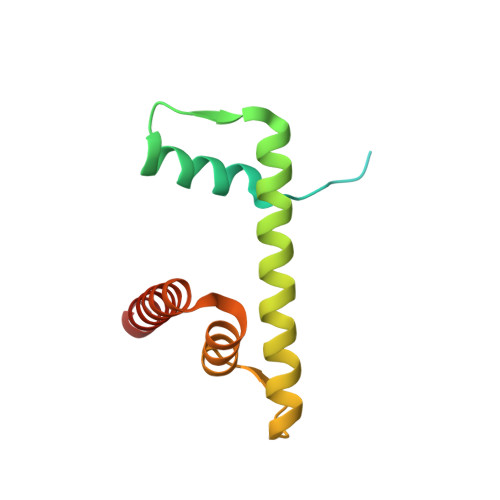
Novel metal(II) arene 2-pyridinecarbothioamides: a rationale to orally active organometallic anticancer agents
Meier, S.M., Hanif, M., Adhireksan, Z., Pichler, V., Novak, M., Jirkovsky, E., Jakupec, M.A., Arion, V.B., Davey, C.A., Keppler, B.K., Hartinger, C.G.(2013) Chem Sci 4: 1837-1846