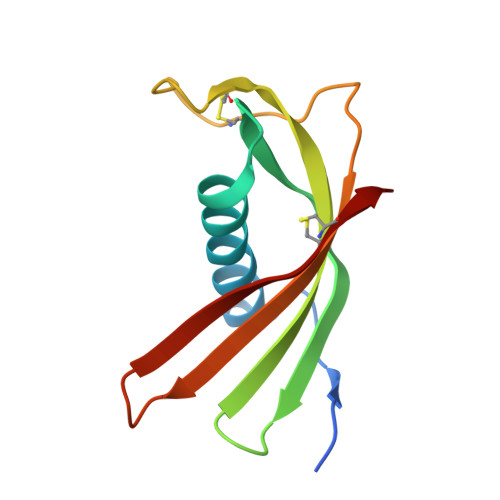
Structural Basis for the Immunomodulatory Function of Cysteine Protease Inhibitor from Human Roundworm Ascaris lumbricoides.
Mei, G., Dong, J., Li, Z., Liu, S., Liu, Y., Sun, M., Liu, G., Su, Z., Liu, J.(2014) PLoS One 9: e96069-e96069
- PubMed: 24781326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0096069
- Primary Citation Related Structures:
4IT7 - PubMed Abstract:
Immunosuppression associated with infections of nematode parasites has been documented. Cysteine protease inhibitor (CPI) released by the nematode parasites is identified as one of the major modulators of host immune response. In this report, we demonstrated that the recombinant CPI protein of Ascaris lumbricoides (Al-CPI) strongly inhibited the activities of cathepsin L, C, S, and showed weaker effect to cathepsin B. Crystal structure of Al-CPI was determined to 2.1 Å resolution. Two segments of Al-CPI, loop 1 and loop 2, were proposed as the key structure motifs responsible for Al-CPI binding with proteases and its inhibitory activity. Mutations at loop 1 and loop 2 abrogated the protease inhibition activity to various extents. These results provide the molecular insight into the interaction between the nematode parasite and its host and will facilitate the development of anthelmintic agents or design of anti-autoimmune disease drugs.
- State Key Laboratory of Respiratory Disease, Guangzhou Institutes of Biomedicine and Health, Chinese Academy of Sciences, Guangzhou, China; School of Life Sciences, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: