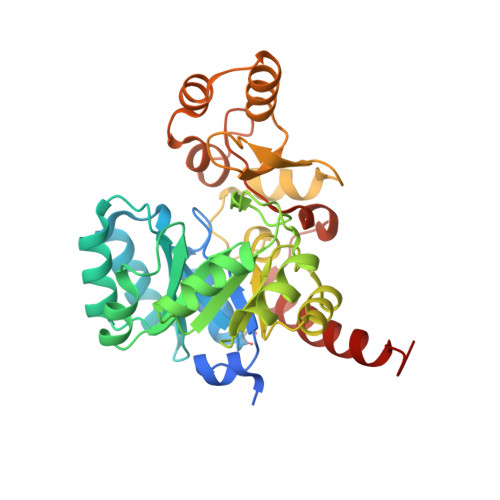
Structural characterization of Porphyromonas gingivalis enoyl-ACP reductase II (FabK).
Hevener, K.E., Santarsiero, B.D., Lee, H., Jones, J.A., Boci, T., Johnson, M.E., Mehboob, S.(2018) Acta Crystallogr F Struct Biol Commun 74: 105-112
- PubMed: 29400320 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18000262
- Primary Citation Related Structures:
4IQL - PubMed Abstract:
Enoyl-acyl carrier protein (ACP) reductase II (FabK) is a critical rate-limiting enzyme in the bacterial type II fatty-acid synthesis (FAS II) pathway. FAS II pathway enzymes are markedly disparate from their mammalian analogs in the FAS I pathway in both structure and mechanism. Enzymes involved in bacterial fatty-acid synthesis represent viable drug targets for Gram-negative pathogens, and historical precedent exists for targeting them in the treatment of diseases of the oral cavity. The Gram-negative organism Porphyromonas gingivalis represents a key causative agent of the costly and highly prevalent disease known as chronic periodontitis, and exclusively expresses FabK as its enoyl reductase enzyme in the FAS-II pathway. Together, these characteristics distinguish P. gingivalis FabK (PgFabK) as an attractive and novel narrow-spectrum antibacterial target candidate. PgFabK is a flavoenzyme that is dependent on FMN and NADPH as cofactors for the enzymatic reaction, which reduces the enoyl substrate via a ping-pong mechanism. Here, the structure of the PgFabK enzyme as determined using X-ray crystallography is reported to 1.9 Å resolution with endogenous FMN fully resolved and the NADPH cofactor partially resolved. PgFabK possesses a TIM-barrel motif, and all flexible loops are visible. The determined structure has allowed insight into the structural basis for the NADPH dependence observed in PgFabK and the role of a monovalent cation that has been observed in previous studies to be stringently required for FabK activity. The PgFabK structure and the insights gleaned from its analysis will facilitate structure-based drug-discovery efforts towards the prevention and treatment of P. gingivalis infection.
- Department of Pharmaceutical Sciences, University of Tennessee Health Science Center, 881 Madison Avenue, Suite 571, Memphis, TN 38163-2198, USA.
Organizational Affiliation: