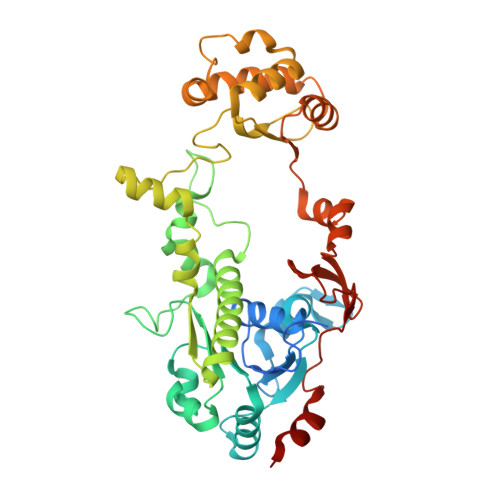
Function and X-Ray crystal structure of Escherichia coli YfdE.
Mullins, E.A., Sullivan, K.L., Kappock, T.J.(2013) PLoS One 8: e67901-e67901
- PubMed: 23935849 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0067901
- Primary Citation Related Structures:
4HL6 - PubMed Abstract:
Many food plants accumulate oxalate, which humans absorb but do not metabolize, leading to the formation of urinary stones. The commensal bacterium Oxalobacter formigenes consumes oxalate by converting it to oxalyl-CoA, which is decarboxylated by oxalyl-CoA decarboxylase (OXC). OXC and the class III CoA-transferase formyl-CoA:oxalate CoA-transferase (FCOCT) are widespread among bacteria, including many that have no apparent ability to degrade or to resist external oxalate. The EvgA acid response regulator activates transcription of the Escherichia coli yfdXWUVE operon encoding YfdW (FCOCT), YfdU (OXC), and YfdE, a class III CoA-transferase that is ~30% identical to YfdW. YfdW and YfdU are necessary and sufficient for oxalate-induced protection against a subsequent acid challenge; neither of the other genes has a known function. We report the purification, in vitro characterization, 2.1-Å crystal structure, and functional assignment of YfdE. YfdE and UctC, an orthologue from the obligate aerobe Acetobacter aceti, perform the reversible conversion of acetyl-CoA and oxalate to oxalyl-CoA and acetate. The annotation of YfdE as acetyl-CoA:oxalate CoA-transferase (ACOCT) expands the scope of metabolic pathways linked to oxalate catabolism and the oxalate-induced acid tolerance response. FCOCT and ACOCT active sites contain distinctive, conserved active site loops (the glycine-rich loop and the GNxH loop, respectively) that appear to encode substrate specificity.
- Department of Biochemistry, Purdue University, West Lafayette, Indiana, United States of America.
Organizational Affiliation: