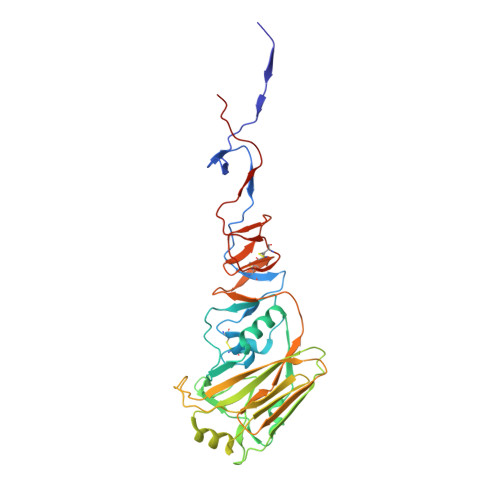
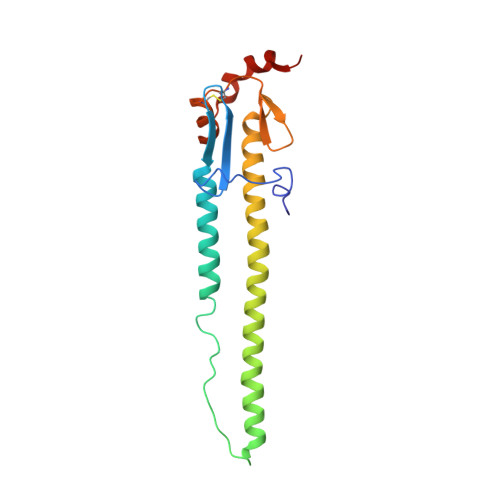
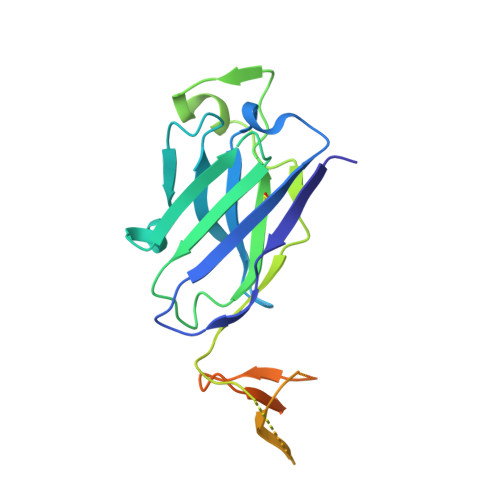
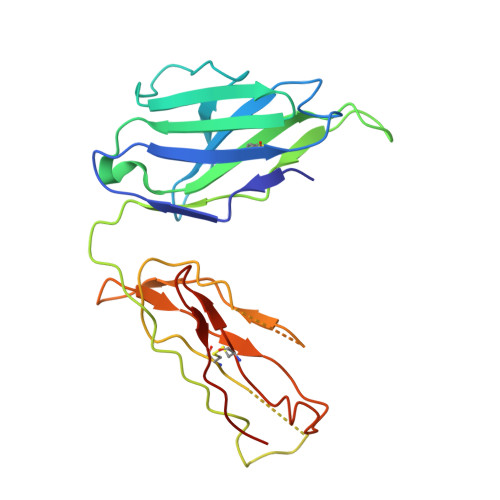
A recurring motif for antibody recognition of the receptor-binding site of influenza hemagglutinin.
Xu, R., Krause, J.C., McBride, R., Paulson, J.C., Crowe, J.E., Wilson, I.A.(2013) Nat Struct Mol Biol 20: 363-370
- PubMed: 23396351 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2500
- Primary Citation Related Structures:
4HF5, 4HFU, 4HG4 - PubMed Abstract:
Influenza virus hemagglutinin (HA) mediates receptor binding and viral entry during influenza infection. The development of receptor analogs as viral-entry blockers has not been successful, which suggests that sialic acid may not be an ideal scaffold to obtain broad, potent HA inhibitors. Here, we report crystal structures of Fab fragments from three human antibodies that neutralize the 1957 pandemic H2N2 influenza virus in complex with H2 HA. All three antibodies use an aromatic residue to plug a conserved cavity in the HA receptor-binding site. Each antibody interacts with the absolutely conserved HA1 Trp153 at the cavity base through π-π stacking with the signature Phe54 of two VH1-69-encoded antibodies or a tyrosine from HCDR3 in the other antibody. This highly conserved interaction can be used as a starting point to design inhibitors targeting this conserved hydrophobic pocket in influenza viruses.
- Department of Integrative Structural and Computational Biology, Scripps Research Institute, La Jolla, California, USA.
Organizational Affiliation: