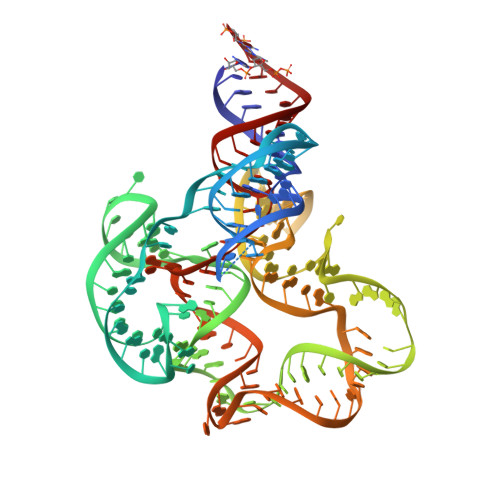
Structural insights into ligand binding and gene expression control by an adenosylcobalamin riboswitch.
Peselis, A., Serganov, A.(2012) Nat Struct Mol Biol 19: 1182-1184
- PubMed: 23064646 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2405
- Primary Citation Related Structures:
4GXY - PubMed Abstract:
Coenzyme B(12) has a key role in various enzymatic reactions and controls expression of bacterial genes through riboswitches. Here we report the crystal structure of the Symbiobacterium thermophilum B(12) riboswitch bound to its ligand adenosylcobalamin. The riboswitch forms a unique junctional structure with a large ligand-binding pocket tailored for specific recognition of the adenosyl moiety and flanked by structural elements that stabilize the regulatory region and enable control of gene expression.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, New York, USA.
Organizational Affiliation: