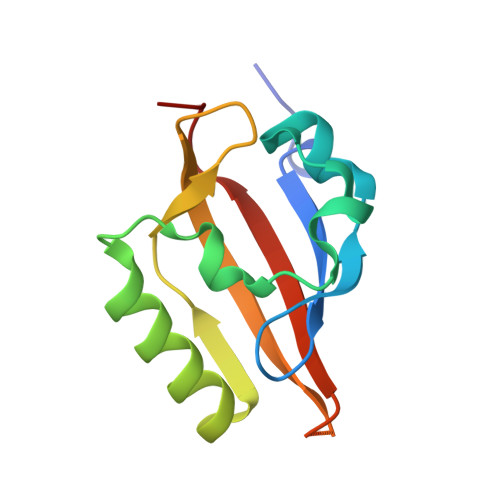
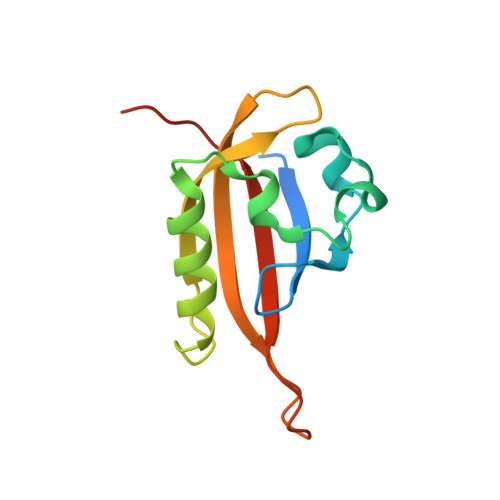
Development of Inhibitors of the PAS-B Domain of the HIF-2 alpha Transcription Factor
Rogers, J.L., Bayeh, L., Scheuermann, T.H., Longgood, J., Key, J., Naidoo, J., Melito, L., Shokri, C., Frantz, D.E., Bruick, R.K., Gardner, K.H., Macmillan, J.B., Tambar, U.K.(2013) J Med Chem 56: 1739-1747
- PubMed: 23363003 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm301847z
- Primary Citation Related Structures:
4GS9 - PubMed Abstract:
Hypoxia inducible factors (HIFs) are heterodimeric transcription factors induced in a variety of pathophysiological settings, including cancer. We describe the first detailed structure-activity relationship study of small molecules designed to inhibit HIF-2α-ARNT heterodimerization by binding an internal cavity of the HIF-2α PAS-B domain. Through a series of biophysical characterizations of inhibitor-protein interactions (NMR and X-ray crystallography), we have established the structural requirements for artificial inhibitors of the HIF-2α-ARNT PAS-B interaction. These results may serve as a foundation for discovering therapeutic agents that function by a novel mode of action.
- Department of Biochemistry, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, Texas 75390-9038, United States.
Organizational Affiliation: