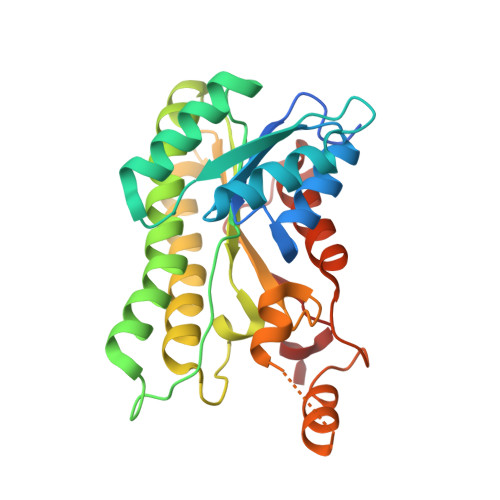
Crystal structures of S-HPCDH reveal determinants of stereospecificity for R- and S-hydroxypropyl-coenzyme M dehydrogenases.
Bakelar, J.W., Sliwa, D.A., Johnson, S.J.(2013) Arch Biochem Biophys 533: 62-68
- PubMed: 23474457 Search on PubMed
- DOI: https://doi.org/10.1016/j.abb.2013.02.017
- Primary Citation Related Structures:
4GH5, 4ITU - PubMed Abstract:
(R)- and (S)-hydroxypropyl-coenzyme M dehydrogenases (R- and S-HPCDH) are stereospecific enzymes that are central to the metabolism of propylene and epoxide in Xanthobacter autotrophicus. The bacterium produces R- and S-HPCDH simultaneously to facilitate transformation of R- and S-enantiomers of epoxypropane to a common achiral product 2-ketopropyl-CoM (2-KPC). Both R- and S-HPCDH are highly specific for their respective substrates as each enzyme displays less than 0.5% activity with the opposite substrate isomer. In order to elucidate the structural basis for stereospecificity displayed by R- and S-HPCDH we have determined substrate bound crystal structures of S-HPCDH to 1.6Å resolution. Comparisons to the previously reported product-bound structure of R-HPCDH reveal that although the placement of catalytic residues within the active site of each enzyme is nearly identical, structural differences in the surrounding area provide each enzyme with a distinct substrate binding pocket. These structures demonstrate how chiral discrimination by R- and S-HPCDH results from alternative binding of the distal end of substrates within each substrate binding pocket.
- Department of Chemistry & Biochemistry, Utah State University, Logan, UT 84322-0300, USA.
Organizational Affiliation: