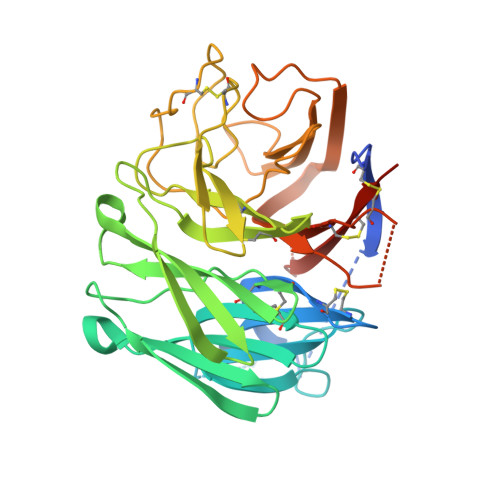
Crystal structures of two subtype N10 neuraminidase-like proteins from bat influenza A viruses reveal a diverged putative active site.
Zhu, X., Yang, H., Guo, Z., Yu, W., Carney, P.J., Li, Y., Chen, L.M., Paulson, J.C., Donis, R.O., Tong, S., Stevens, J., Wilson, I.A.(2012) Proc Natl Acad Sci U S A 109: 18903-18908
- PubMed: 23012478 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1212579109
- Primary Citation Related Structures:
4GDI, 4GDJ, 4GEZ - PubMed Abstract:
Recently, we reported a unique influenza A virus subtype H17N10 from little yellow-shouldered bats. Its neuraminidase (NA) gene encodes a protein that appears to be highly divergent from all known influenza NAs and was assigned as a new subtype N10. To provide structural and functional insights on the bat H17N10 virus, X-ray structures were determined for N10 NA proteins from influenza A viruses A/little yellow-shouldered bat/Guatemala/164/2009 (GU09-164) in two crystal forms at 1.95 Å and 2.5 Å resolution and A/little yellow-shouldered bat/Guatemala/060/2010 (GU10-060) at 2.0 Å. The overall N10 structures are similar to each other and to other known influenza NA structures, with a single highly conserved calcium binding site in each monomer. However, the region corresponding to the highly conserved active site of influenza A N1-N9 NA subtypes and influenza B NA differs substantially. In particular, most of the amino acid residues required for NA activity are substituted, and the putative active site is much wider because of displacement of the 150-loop and 430-loop. These structural features and the fact that the recombinant N10 protein exhibits no, or extremely low, NA activity suggest that it may have a different function than the NA proteins of other influenza viruses. Accordingly, we propose that the N10 protein be termed an NA-like protein until its function is elucidated.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: