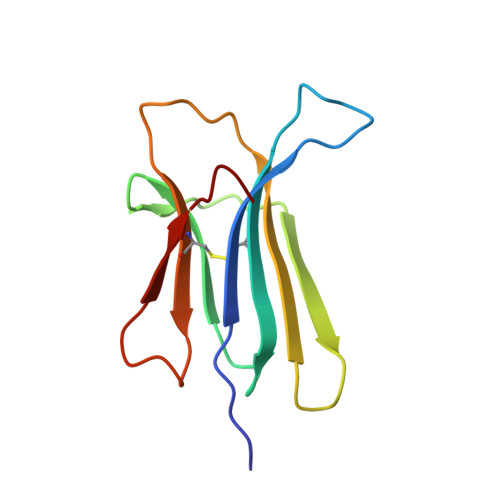
Hereditary systemic amyloidosis due to Asp76Asn variant beta-2-microglobulin.
Valleix, S., Gillmore, J.D., Bridoux, F., Mangione, P.P., Dogan, A., Nedelec, B., Boimard, M., Touchard, G., Goujon, J.M., Lacombe, C., Lozeron, P., Adams, D., Lacroix, C., Maisonobe, T., Plante-Bordeneuve, V., Vrana, J.A., Theis, J.D., Giorgetti, S., Porcari, R., Ricagno, S., Bolognesi, M., Stoppini, M., Delpech, M., Pepys, M.B., Hawkins, P.N., Bellotti, V.(2012) N Engl J Med 366: 2276-2283
- PubMed: 22693999 Search on PubMed
- DOI: https://doi.org/10.1056/NEJMoa1201356
- Primary Citation Related Structures:
4FXL - PubMed Abstract:
We describe a kindred with slowly progressive gastrointestinal symptoms and autonomic neuropathy caused by autosomal dominant, hereditary systemic amyloidosis. The amyloid consists of Asp76Asn variant β(2)-microglobulin. Unlike patients with dialysis-related amyloidosis caused by sustained high plasma concentrations of wild-type β(2)-microglobulin, the affected members of this kindred had normal renal function and normal circulating β(2)-microglobulin values. The Asp76Asn β(2)-microglobulin variant was thermodynamically unstable and remarkably fibrillogenic in vitro under physiological conditions. Previous studies of β(2)-microglobulin aggregation have not shown such amyloidogenicity for single-residue substitutions. Comprehensive biophysical characterization of the β(2)-microglobulin variant, including its 1.40-Å, three-dimensional structure, should allow further elucidation of fibrillogenesis and protein misfolding.
- Laboratoire de Biochimie et de Génétique Moléculaire, Université Paris-Descartes, Sorbonne Paris Cité, Faculté de Médecine Paris, Assistance Public–Hôpitaux de Paris (AP-HP), Paris, France. sophie.valleix@cch.aphp.fr
Organizational Affiliation: