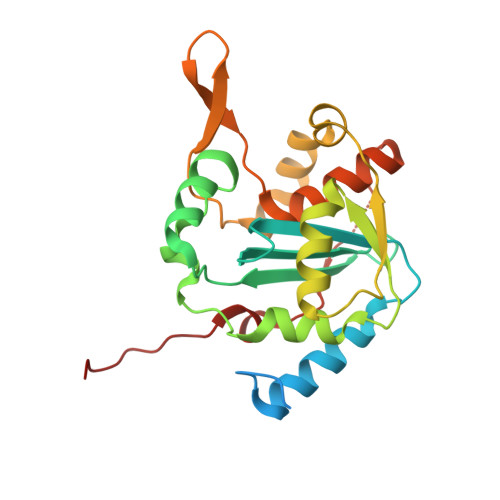
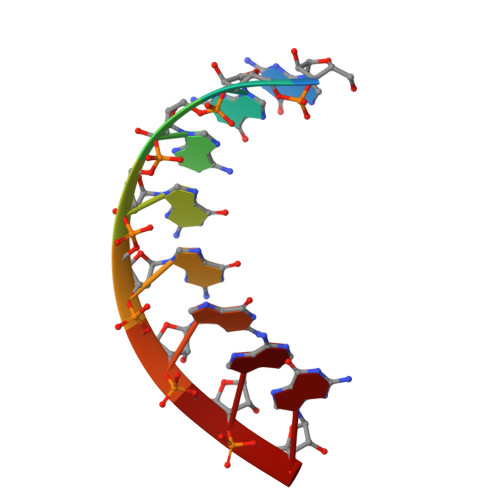
Structural Basis for the dsRNA Specificity of the Lassa Virus NP Exonuclease.
Hastie, K.M., King, L.B., Zandonatti, M.A., Saphire, E.O.(2012) PLoS One 7: e44211-e44211
- PubMed: 22937163 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0044211
- Primary Citation Related Structures:
4FVU - PubMed Abstract:
Lassa virus causes hemorrhagic fever characterized by immunosuppression. The nucleoprotein of Lassa virus, termed NP, binds the viral genome. It also has an additional enzymatic activity as an exonuclease that specifically digests double-stranded RNA (dsRNA). dsRNA is a strong signal to the innate immune system of viral infection. Digestion of dsRNA by the NP exonuclease activity appears to cause suppression of innate immune signaling in the infected cell. Although the fold of the NP enzyme is conserved and the active site completely conserved with other exonucleases in its DEDDh family, NP is atypical among exonucleases in its preference for dsRNA and its strict specificity for one substrate. Here, we present the crystal structure of Lassa virus NP in complex with dsRNA. We find that unlike the exonuclease in Klenow fragment, the double-stranded nucleic acid in complex with Lassa NP remains base-paired instead of splitting, and that binding of the paired complementary strand is achieved by "relocation" of a basic loop motif from its typical exonuclease position. Further, we find that just one single glycine that contacts the substrate strand and one single tyrosine that stacks with a base of the complementary, non-substrate strand are responsible for the unique substrate specificity. This work thus provides templates for development of antiviral drugs that would be specific for viral, rather than host exonucleases of similar fold and active site, and illustrates how a very few amino acid changes confer alternate specificity and biological phenotype to an enzyme.
- Department of Immunology and Microbial Science, The Scripps Research Institute, La Jolla, California, United States of America.
Organizational Affiliation: