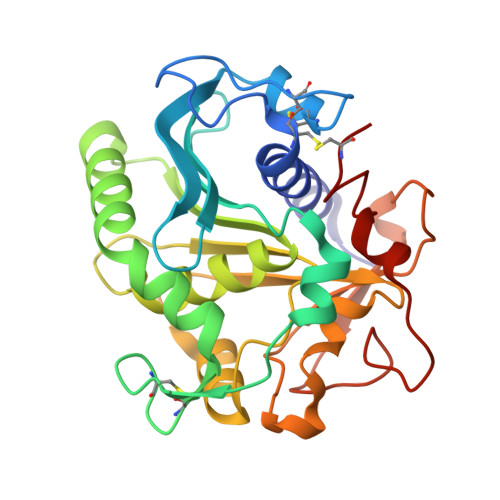
Enhancement of stability of a lipase by subjecting to three phase partitioning (TPP): structures of native and TPP-treated lipase from Thermomyces lanuginosa
Kumar, M., Mukherjee, J., Sinha, M., Kaur, P., Sharma, S., Gupta, M.N., Singh, T.P.(2015) Sustain Chem Process