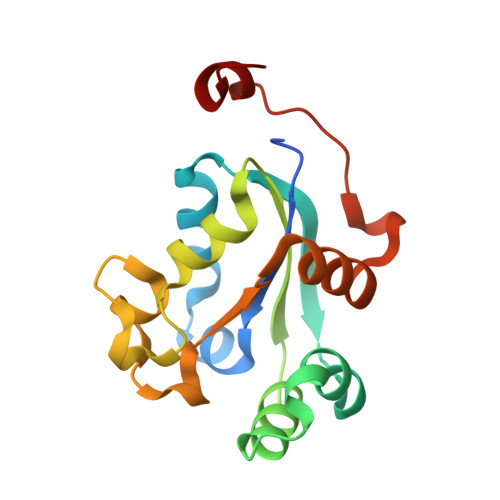
Structural and Functional Analyses of Trypanosoma brucei Nucleoside Diphosphate Kinase
Makori, P., Boeckman, M.P., David, H.S., Payne, F., Gatling, M., Greer, C., Hayes, D., Jefferson, A., Maxwell, M., Smith, C., Watson, J., Williams, L., Barkley, J., Pepper, C., Zininga, T., Subramanian, S., Abramov, A., Gardberg, A.S., Edwards, T.E., Staker, B.L., Stewart, L.J., Myler, P.J., Asojo, O.A., Jeje, O., Fanucchi, S., Achilonu, I., Smith, C.L., Chakafana, G.(2026) ACS Omega