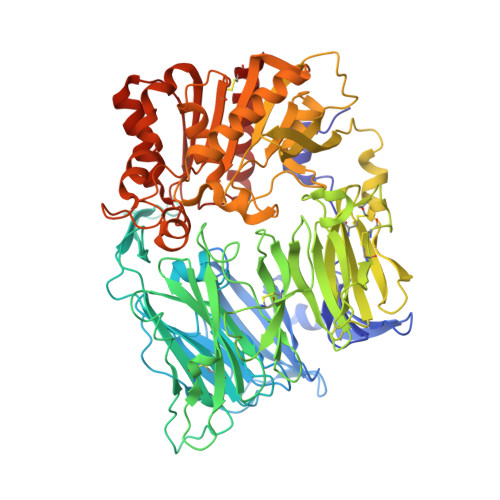
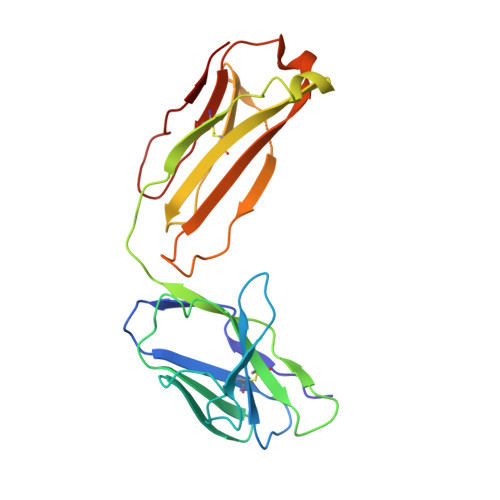
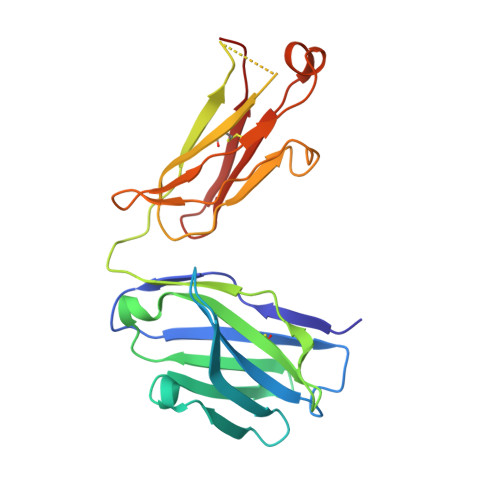
An Inhibitory Antibody Against DPP IV Improves Glucose Tolerance in vivo - Validation of Large Molecule Approach for DPP IV Inhibition
Tang, J., Majeti, J., Sudom, A., Xiong, Y., Lu, M., Liu, Q., Higbee, J., Zhang, Y., Wang, Y., Wang, W., Cao, P., Xia, Z., Johnstone, S., Yang, X., Yu, T., Sharkov, N., Walker, N., Tu, H., Shen, W., Wang, Z.To be published.