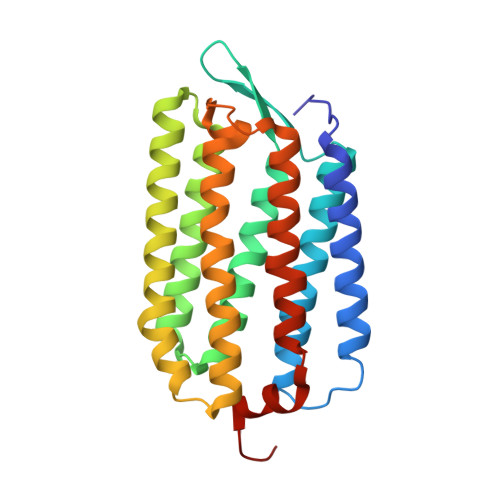
Crystal structure of deltarhodopsin-3 from Haloterrigena thermotolerans
Zhang, J., Mizuno, K., Murata, Y., Koide, H., Murakami, M., Ihara, K., Kouyama, T.(2013) Proteins 81: 1585-1592
- PubMed: 23625688 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24316
- Primary Citation Related Structures:
4FBZ - PubMed Abstract:
Deltarhodopsin, a new member of the microbial rhodopsin family, functions as a light-driven proton pump. Here, we report the three-dimensional structure of deltarhodopsin (dR3) from Haloterrigena thermotolerans at 2.7 Å resolution. A crystal belonging to space group R32 (a, b = 111.71 Å, c = 198.25 Å) was obtained by the membrane fusion method. In this crystal, dR3 forms a trimeric structure as observed for bacteriorhodopsin (bR). Structural comparison of dR with bR showed that the inner part (the proton release and uptake pathways) is highly conserved. Meanwhile, residues in the protein-protein contact region are largely altered so that the diameter of the trimeric structure at the cytoplasmic side is noticeably larger in dR3. Unlike bR, dR3 possesses a helical segment at the C-terminal region that fills the space between the AB and EF loops. A significant difference is also seen in the FG loop, which is one residue longer in dR3. Another peculiar property of dR3 is a highly crowded distribution of positively charged residues on the cytoplasmic surface, which may be relevant to a specific interaction with some cytoplasmic component.
- Department of Physics, Graduate School of Science, Nagoya University, Nagoya, Japan.
Organizational Affiliation: