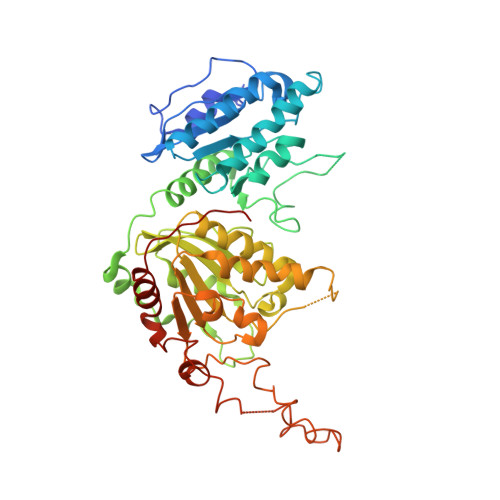
Structure, Activity, and Inhibition of the Carboxyltransferase beta-Subunit of Acetyl Coenzyme A Carboxylase (AccD6) from Mycobacterium tuberculosis.
Reddy, M.C., Breda, A., Bruning, J.B., Sherekar, M., Valluru, S., Thurman, C., Ehrenfeld, H., Sacchettini, J.C.(2014) Antimicrob Agents Chemother 58: 6122-6132
- PubMed: 25092705 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.02574-13
- Primary Citation Related Structures:
4FB8, 4G2R - PubMed Abstract:
In Mycobacterium tuberculosis, the carboxylation of acetyl coenzyme A (acetyl-CoA) to produce malonyl-CoA, a building block in long-chain fatty acid biosynthesis, is catalyzed by two enzymes working sequentially: a biotin carboxylase (AccA) and a carboxyltransferase (AccD). While the exact roles of the three different biotin carboxylases (AccA1 to -3) and the six carboxyltransferases (AccD1 to -6) in M. tuberculosis are still not clear, AccD6 in complex with AccA3 can synthesize malonyl-CoA from acetyl-CoA. A series of 10 herbicides that target plant acetyl-CoA carboxylases (ACC) were tested for inhibition of AccD6 and for whole-cell activity against M. tuberculosis. From the tested herbicides, haloxyfop, an arylophenoxypropionate, showed in vitro inhibition of M. tuberculosis AccD6, with a 50% inhibitory concentration (IC50) of 21.4 ± 1 μM. Here, we report the crystal structures of M. tuberculosis AccD6 in the apo form (3.0 Å) and in complex with haloxyfop-R (2.3 Å). The structure of M. tuberculosis AccD6 in complex with haloxyfop-R shows two molecules of the inhibitor bound on each AccD6 subunit. These results indicate the potential for developing novel therapeutics for tuberculosis based on herbicides with low human toxicity.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, Texas, USA.
Organizational Affiliation: