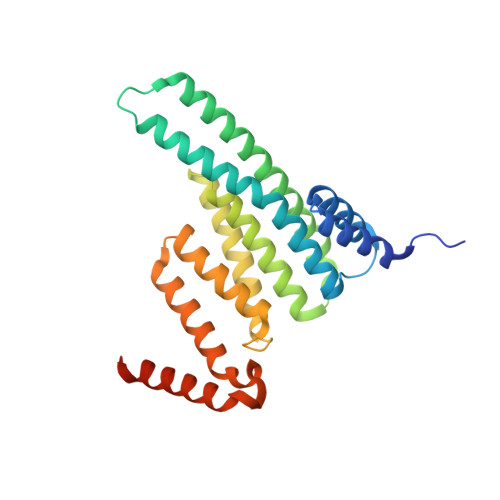
The Crystal Structure of Giardia duodenalis 14-3-3 in the Apo Form: When Protein Post-Translational Modifications Make the Difference.
Fiorillo, A., di Marino, D., Bertuccini, L., Via, A., Pozio, E., Camerini, S., Ilari, A., Lalle, M.(2014) PLoS One 9: e92902-e92902
- PubMed: 24658679 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0092902
- Primary Citation Related Structures:
4F7R - PubMed Abstract:
The 14-3-3s are a family of dimeric evolutionary conserved pSer/pThr binding proteins that play a key role in multiple biological processes by interacting with a plethora of client proteins. Giardia duodenalis is a flagellated protozoan that affects millions of people worldwide causing an acute and chronic diarrheal disease. The single giardial 14-3-3 isoform (g14-3-3), unique in the 14-3-3 family, needs the constitutive phosphorylation of Thr214 and the polyglycylation of its C-terminus to be fully functional in vivo. Alteration of the phosphorylation and polyglycylation status affects the parasite differentiation into the cyst stage. To further investigate the role of these post-translational modifications, the crystal structure of the g14-3-3 was solved in the unmodified apo form. Oligomers of g14-3-3 were observed due to domain swapping events at the protein C-terminus. The formation of filaments was supported by TEM. Mutational analysis, in combination with native PAGE and chemical cross-linking, proved that polyglycylation prevents oligomerization. In silico phosphorylation and molecular dynamics simulations supported a structural role for the phosphorylation of Thr214 in promoting target binding. Our findings highlight unique structural features of g14-3-3 opening novel perspectives on the evolutionary history of this protein family and envisaging the possibility to develop anti-giardial drugs targeting g14-3-3.
- Department of Biochemical Sciences "A. Rossi-Fanelli", University of Rome "Sapienza", Rome, Italy; Institute of Molecular Biology and Pathology, CNR, Rome, Italy and Institute Pasteur Cenci-Bolognetti Foundation at Department of Biochemical Sciences "A. Rossi-Fanelli", University of Rome "Sapienza", Rome, Italy.
Organizational Affiliation: