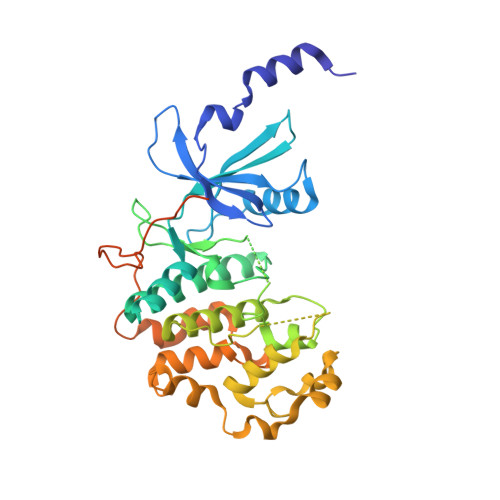
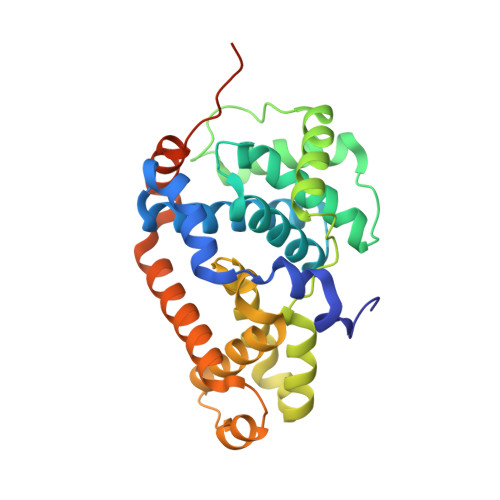
Structure-kinetic relationship study of CDK8/CycC specific compounds.
Schneider, E.V., Bottcher, J., Huber, R., Maskos, K., Neumann, L.(2013) Proc Natl Acad Sci U S A 110: 8081-8086
- PubMed: 23630251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1305378110
- Primary Citation Related Structures:
4F6S, 4F6U, 4F6W, 4F70, 4F7J, 4F7L, 4F7N, 4F7S, 4G6L - PubMed Abstract:
In contrast with the very well explored concept of structure-activity relationship, similar studies are missing for the dependency between binding kinetics and compound structure of a protein ligand complex, the structure-kinetic relationship. Here, we present a structure-kinetic relationship study of the cyclin-dependent kinase 8 (CDK8)/cyclin C (CycC) complex. The scaffold moiety of the compounds is anchored in the kinase deep pocket and extended with diverse functional groups toward the hinge region and the front pocket. These variations can cause the compounds to change from fast to slow binding kinetics, resulting in an improved residence time. The flip of the DFG motif ("DMG" in CDK8) to the inactive DFG-out conformation appears to have relatively little influence on the velocity of binding. Hydrogen bonding with the kinase hinge region contributes to the residence time but has less impact than hydrophobic complementarities within the kinase front pocket.
- Max-Planck-Institut für Biochemie, D-82152 Martinsried, Germany. schneider@proteros.com
Organizational Affiliation: