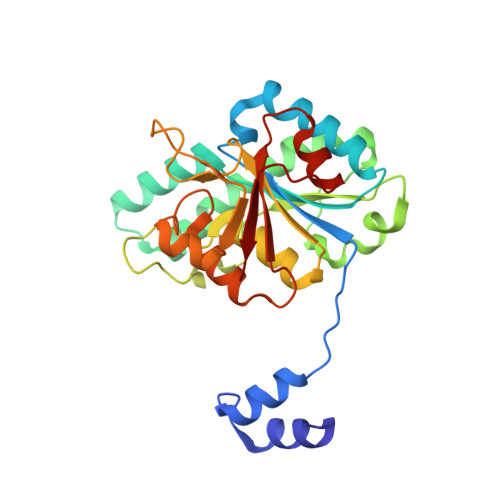
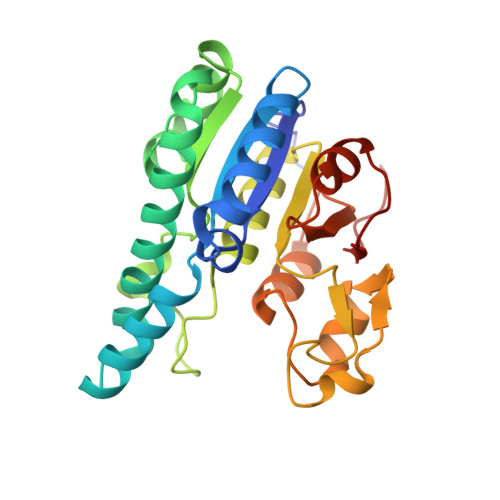
Nature's Polyoxometalate Chemistry: X-ray Structure of the Mo Storage Protein Loaded with Discrete Polynuclear Mo-O Clusters.
Kowalewski, B., Poppe, J., Demmer, U., Warkentin, E., Dierks, T., Ermler, U., Schneider, K.(2012) J Am Chem Soc 134: 9768-9774
- PubMed: 22612644 Search on PubMed
- DOI: https://doi.org/10.1021/ja303084n
- Primary Citation Related Structures:
4F6T - PubMed Abstract:
Some N(2)-fixing bacteria prolong the functionality of nitrogenase in molybdenum starvation by a special Mo storage protein (MoSto) that can store more than 100 Mo atoms. The presented 1.6 Å X-ray structure of MoSto from Azotobacter vinelandii reveals various discrete polyoxomolybdate clusters, three covalently and three noncovalently bound Mo(8), three Mo(5-7), and one Mo(3) clusters, and several low occupied, so far undefinable clusters, which are embedded in specific pockets inside a locked cage-shaped (αβ)(3) protein complex. The structurally identical Mo(8) clusters (three layers of two, four, and two MoO(n) octahedra) are distinguishable from the [Mo(8)O(26)](4-) cluster formed in acidic solutions by two displaced MoO(n) octahedra implicating three kinetically labile terminal ligands. Stabilization in the covalent Mo(8) cluster is achieved by Mo bonding to Hisα156-N(ε2) and Gluα129-O(ε1). The absence of covalent protein interactions in the noncovalent Mo(8) cluster is compensated by a more extended hydrogen-bond network involving three pronounced histidines. One displaced MoO(n) octahedron might serve as nucleation site for an inhomogeneous Mo(5-7) cluster largely surrounded by bulk solvent. In the Mo(3) cluster located on the 3-fold axis, the three accurately positioned His140-N(ε2) atoms of the α subunits coordinate to the Mo atoms. The formed polyoxomolybdate clusters of MoSto, not detectable in bulk solvent, are the result of an interplay between self- and protein-driven assembly processes that unite inorganic supramolecular and protein chemistry in a host-guest system. Template, nucleation/protection, and catalyst functions of the polypeptide as well as perspectives for designing new clusters are discussed.
- Biochemie I, Fakultät für Chemie, Universität Bielefeld, Universitätsstraße 25, D-33615 Bielefeld, Germany.
Organizational Affiliation: