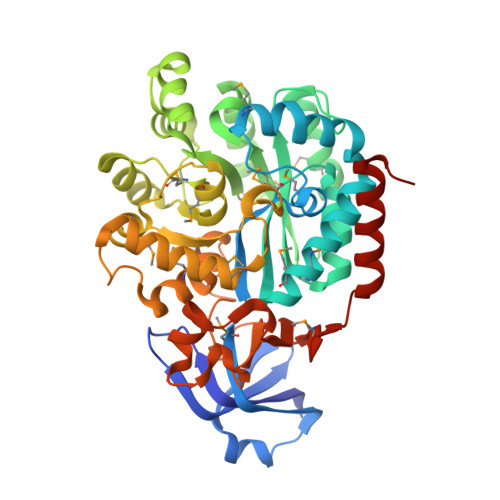
Crystal structure of an adenosine deaminase homolog from Chromobacterium violaceum (target NYSGRC-019589) bound Zn and 5'-Methylthioadenosine (unproductive complex)
Kim, J., Vetting, M.W., Sauder, J.M., Burley, S.K., Raushel, F.M., Bonanno, J.B., Almo, S.C.To be published.