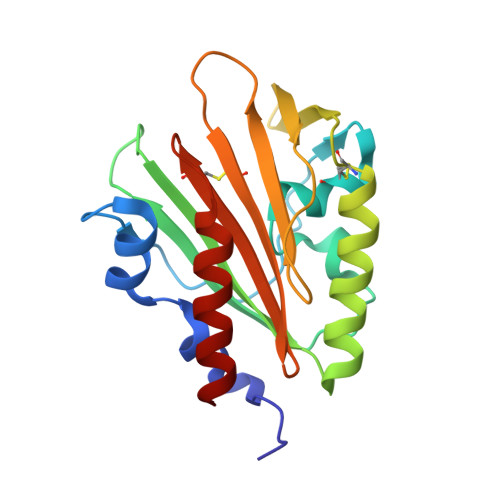
Structure of the sensor domain of Mycobacterium tuberculosis PknH receptor kinase reveals a conserved binding cleft.
Cavazos, A., Prigozhin, D.M., Alber, T.(2012) J Mol Biology 422: 488-494
- PubMed: 22727744 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2012.06.011
- Primary Citation Related Structures:
4ESQ - PubMed Abstract:
Since their discovery over 20 years ago, eukaryotic-like transmembrane receptor Ser/Thr protein kinases (STPKs) have been shown to play critical roles in the virulence, growth, persistence, and reactivation of many bacteria. Information regarding the signals transmitted by these proteins, however, remains scarce. To enhance understanding of the basis for STPK receptor signaling, we determined the 1.7-Å-resolution crystal structure of the extracellular sensor domain of the Mycobacterium tuberculosis receptor STPK, PknH (Rv1266c). The PknH sensor domain adopts an unanticipated fold containing two intramolecular disulfide bonds and a large hydrophobic and polar cleft. The residues lining the cleft and those surrounding the disulfide bonds are conserved. These results suggest that PknH binds a small-molecule ligand that signals by changing the location or quaternary structure of the kinase domain.
- Department of Molecular and Cell Biology, University of California, Berkeley, CA 94720, USA.
Organizational Affiliation: