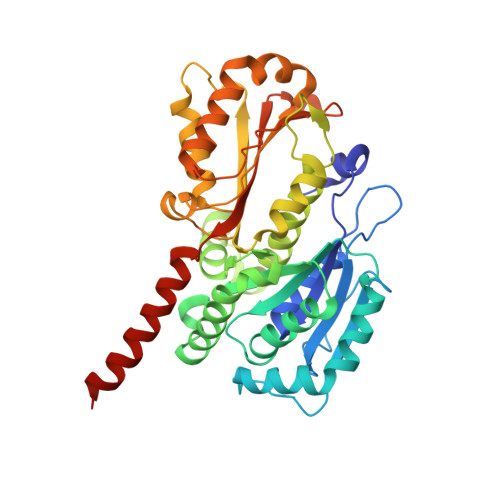
Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid.
Hoshino, S., Hayashi, I.(2012) J Biological Chem 287: 32103-32112
- PubMed: 22847006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.373803
- Primary Citation Related Structures:
4EI7, 4EI8, 4EI9 - PubMed Abstract:
Stable maintenance of low-copy-number plasmids requires partition (par) systems that consist of a nucleotide hydrolase, a DNA-binding protein, and a cis-acting DNA-binding site. The FtsZ/tubulin-like GTPase TubZ was identified as a partitioning factor of the virulence plasmids pBtoxis and pXO1 in Bacillus thuringiensis and Bacillus anthracis, respectively. TubZ exhibits high GTPase activity and assembles into polymers both in vivo and in vitro, and its "treadmilling" movement is required for plasmid stability in the cell. To investigate the molecular mechanism of pXO1 plasmid segregation by TubZ filaments, we determined the crystal structures of Bacillus cereus TubZ in apo-, GDP-, and guanosine 5'-3-O-(thio)triphosphate (GTPγS)-bound forms at resolutions of 2.1, 1.9, and 3.3 Å, respectively. Interestingly, the slowly hydrolyzable GTP analog GTPγS was hydrolyzed to GDP in the crystal. In the post-GTP hydrolysis state, GDP-bound B. cereus TubZ forms a dimer by the head-to-tail association of individual subunits in the asymmetric unit, which is similar to the protofilament formation of FtsZ and B. thuringiensis TubZ. However, the M loop interacts with the nucleotide-binding site of the adjacent subunit and stabilizes the filament structure in a different manner, which indicates that the molecular assembly of the TubZ-related par systems is not stringently conserved. Furthermore, we show that the C-terminal tail of TubZ is required for association with the DNA-binding protein TubR. Using a combination of crystallography, site-directed mutagenesis, and biochemical analysis, our results provide the structural basis of the TubZ polymer that may drive DNA segregation.
- Department of Supramolecular Biology, Graduate School of Nanobioscience, Yokohama City University, 1-7-29 Suehiro, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: