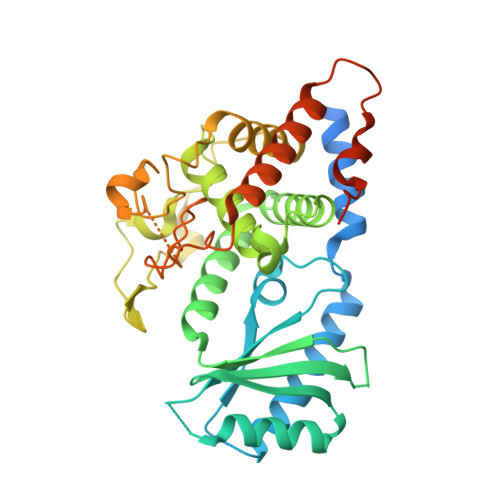
Structural basis for the activity of a cytoplasmic RNA terminal uridylyl transferase.
Yates, L.A., Fleurdepine, S., Rissland, O.S., De Colibus, L., Harlos, K., Norbury, C.J., Gilbert, R.J.(2012) Nat Struct Mol Biol 19: 782-787
- PubMed: 22751018 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2329
- Primary Citation Related Structures:
4E7X, 4E80, 4E8F - PubMed Abstract:
Cytoplasmic terminal uridylyl transferases comprise a conserved family of enzymes that negatively regulate the stability or biological activity of a variety of eukaryotic RNAs, including mRNAs and tumor-suppressor let-7 microRNAs. Here we describe crystal structures of the Schizosaccharomyces pombe cytoplasmic terminal uridylyl transferase Cid1 in two apo conformers and bound to UTP. We demonstrate that a single histidine residue, conserved in mammalian Cid1 orthologs, is responsible for discrimination between UTP and ATP. We also describe a new high-affinity RNA substrate-binding mechanism of Cid1, which is essential for enzymatic activity and is mediated by three basic patches across the surface of the enzyme. Overall, our structures provide a basis for understanding the activity of Cid1 and a mechanism of UTP selectivity conserved in its human orthologs, suggesting potential implications for anticancer drug design.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford OX3 7BN, UK.
Organizational Affiliation: