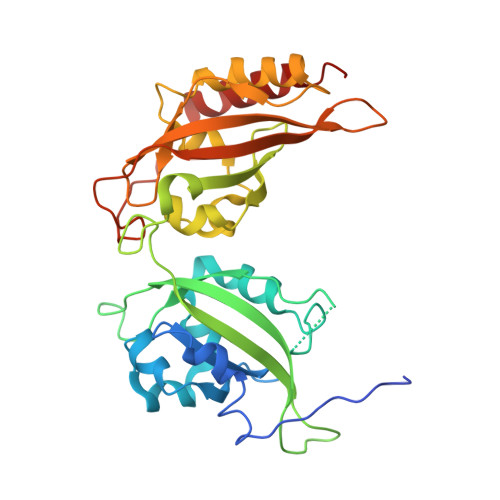
Unwinding the differences of the mammalian PERIOD clock proteins from crystal structure to cellular function.
Kucera, N., Schmalen, I., Hennig, S., Ollinger, R., Strauss, H.M., Grudziecki, A., Wieczorek, C., Kramer, A., Wolf, E.(2012) Proc Natl Acad Sci U S A 109: 3311-3316
- PubMed: 22331899 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1113280109
- Primary Citation Related Structures:
4DJ2, 4DJ3 - PubMed Abstract:
The three PERIOD homologues mPER1, mPER2, and mPER3 constitute central components of the mammalian circadian clock. They contain two PAS (PER-ARNT-SIM) domains (PAS-A and PAS-B), which mediate homo- and heterodimeric mPER-mPER interactions as well as interactions with transcription factors and kinases. Here we present crystal structures of PAS domain fragments of mPER1 and mPER3 and compare them with the previously reported mPER2 structure. The structures reveal homodimers, which are mediated by interactions of the PAS-B β-sheet surface including a highly conserved tryptophan (Trp448(mPER1), Trp419(mPER2), Trp359(mPER3)). mPER1 homodimers are additionally stabilized by interactions between the PAS-A domains and mPER3 homodimers by an N-terminal region including a predicted helix-loop-helix motive. We have verified the existence of these homodimer interfaces in solution and inside cells using analytical gel filtration and luciferase complementation assays and quantified their contributions to homodimer stability by analytical ultracentrifugation. We also show by fluorescence recovery after photobleaching analyses that destabilization of the PAS-B/tryptophan dimer interface leads to a faster mobility of mPER2 containing complexes in human U2OS cells. Our study reveals structural and quantitative differences between the homodimeric interactions of the three mouse PERIOD homologues, which are likely to contribute to their distinct clock functions.
- Max Planck Institute of Molecular Physiology, Department of Structural Biology, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.
Organizational Affiliation: