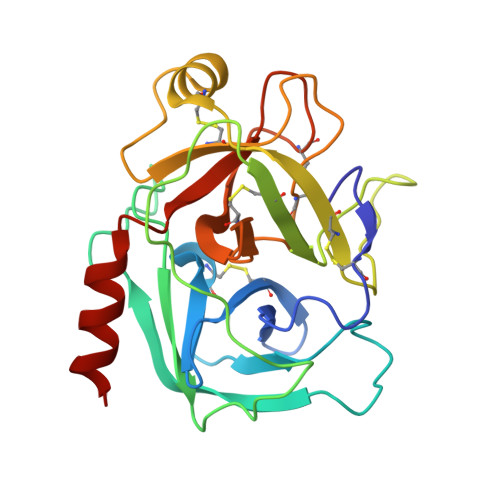
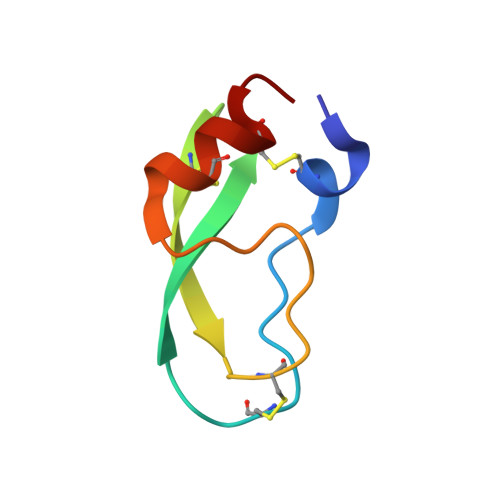
Presence versus absence of hydrogen bond donor Tyr-39 influences interactions of cationic trypsin and mesotrypsin with protein protease inhibitors.
Salameh, M.A., Soares, A.S., Alloy, A., Radisky, E.S.(2012) Protein Sci 21: 1103-1112
- PubMed: 22610453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2097
- Primary Citation Related Structures:
4DG4 - PubMed Abstract:
Mesotrypsin displays unusual resistance to inhibition by polypeptide trypsin inhibitors and cleaves some such inhibitors as substrates, despite a high degree of conservation with other mammalian trypsins. Substitution of Arg for the generally conserved Gly-193 has been implicated as a critical determinant of the unusual behavior of mesotrypsin toward protein protease inhibitors. Another relatively conserved residue near the trypsin active site, Tyr-39, is substituted by Ser-39 in mesotrypsin. Tyr-39, but not Ser-39, forms a hydrogen bond with the main chain amide nitrogen of the P(4) ' residue of a bound protease inhibitor. To investigate the role of the Tyr-39 H-bond in trypsin-inhibitor interactions, we reciprocally mutated position 39 in mesotrypsin and human cationic trypsin to Tyr-39 and Ser-39, respectively. We assessed inhibition constants and cleavage rates of canonical protease inhibitors bovine pancreatic trypsin inhibitor (BPTI) and the amyloid precursor protein Kunitz protease inhibitor domain by mesotrypsin and cationic trypsin variants, finding that the presence of Ser-39 relative to Tyr-39 results in a 4- to 13-fold poorer binding affinity and a 2- to 18-fold increase in cleavage rate. We also report the crystal structure of the mesotrypsin-S39Y•BPTI complex, in which we observe an H-bond between Tyr-39 OH and BPTI Ile-19 N. Our results indicate that the presence of Ser-39 in mesotrypsin, and corresponding absence of a single H-bond to the inhibitor backbone, makes a small but significant functional contribution to the resistance of mesotrypsin to inhibition and the ability of mesotrypsin to proteolyze inhibitors.
- Department of Cancer Biology, Mayo Clinic Cancer Center, Jacksonville, Florida 32224, USA.
Organizational Affiliation: