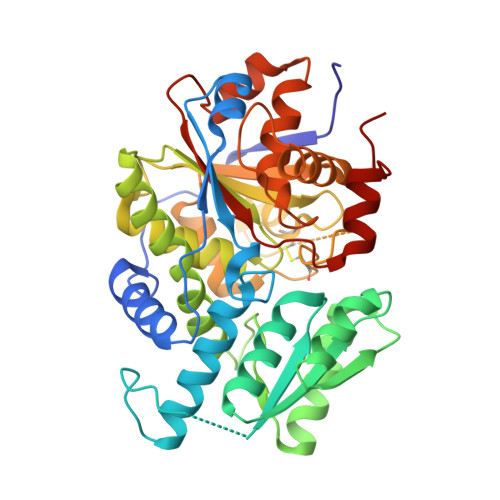
Crystal Structure of Escherichia coli Diaminopropionate Ammonia-lyase Reveals Mechanism of Enzyme Activation and Catalysis
Bisht, S., Rajaram, V., Bharath, S.R., Kalyani, J.N., Khan, F., Rao, A.N., Savithri, H.S., Murthy, M.R.N.(2012) J Biological Chem 287: 20369-20381
- PubMed: 22505717 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.351809
- Primary Citation Related Structures:
4D9G, 4D9I, 4D9K, 4D9M, 4D9N - PubMed Abstract:
Pyridoxal 5'-phosphate (PLP)-dependent enzymes utilize the unique chemistry of a pyridine ring to carry out diverse reactions involving amino acids. Diaminopropionate (DAP) ammonia-lyase (DAPAL) is a prokaryotic PLP-dependent enzyme that catalyzes the degradation of d- and l-forms of DAP to pyruvate and ammonia. Here, we report the first crystal structure of DAPAL from Escherichia coli (EcDAPAL) in tetragonal and monoclinic forms at 2.0 and 2.2 Å resolutions, respectively. Structures of EcDAPAL soaked with substrates were also determined. EcDAPAL has a typical fold type II PLP-dependent enzyme topology consisting of a large and a small domain with the active site at the interface of the two domains. The enzyme is a homodimer with a unique biological interface not observed earlier. Structure of the enzyme in the tetragonal form had PLP bound at the active site, whereas the monoclinic structure was in the apo-form. Analysis of the apo and holo structures revealed that the region around the active site undergoes transition from a disordered to ordered state and assumes a conformation suitable for catalysis only upon PLP binding. A novel disulfide was found to occur near a channel that is likely to regulate entry of ligands to the active site. EcDAPAL soaked with dl-DAP revealed density at the active site appropriate for the reaction intermediate aminoacrylate, which is consistent with the observation that EcDAPAL has low activity under crystallization conditions. Based on the analysis of the structure and results of site-directed mutagenesis, a two-base mechanism of catalysis involving Asp(120) and Lys(77) is suggested.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.
Organizational Affiliation: